1
5-mC | 5-methylcystosine (monclonal, clone 33D3)
- Product Info
-
Sub class: IgG1 Immunogen: OVA-conjugated molecule: 5-methylcytosine (5-mC) Host: Mouse Clonality: Monoclonal Purity: Protein A purified. Format: Liquid Quantity: 100 �g Storage: Store at -80�C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Dot blot (Dot), ELISA (ELISA), Immunofluorescence (IF), MeDIP/MeDIP-seq (methylated DNA Immunoprecipitation (IP) Recommended dilution: 1 : 250 (Dot), 1 : 100 (IF), 0,5-1 �g (MeDIP/MeDIP-seq) - Reactivity
-
Confirmed reactivity: Human Predicted reactivity: Mouse, broad species range Not reactive in: No confirmed exceptions from predicted reactivity are currently known - Application Examples
-
application example 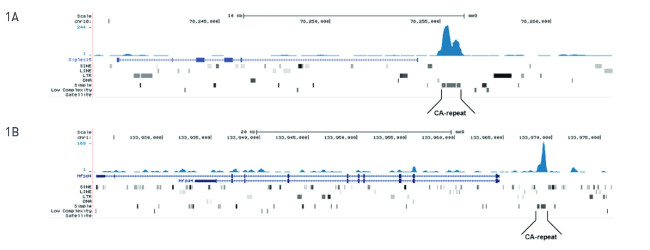
MeDIP-seq: Genomic DNA from E14 ES cells was sheared with the Bioruptor ® to generate random fragments (size range 300 to 700 bp). One μg of the fragmented DNA was ligated to Illumina adapters and the resulting DNA was used for a standard MeDIP assay, using 2 μg of monoclonal against 5-mC. After recovery of the methylated DNA, Illumina sequencing libraries were generated and sequenced on an Illumina Genome Analyzer according to the manufacturer’s instructions. Figure 1A and 1B show Genome browser views of CA simple repeat elements with read distributions specific for 5-mC at 2 gene locations (SigleC15 and Mfsd4). Visual inspection of the peak profiles in a genome browser reveals high enrichment of CA simple repeats in affinity-enriched methylated fragments after MeDIP with the 5-mC monoclonal antibody.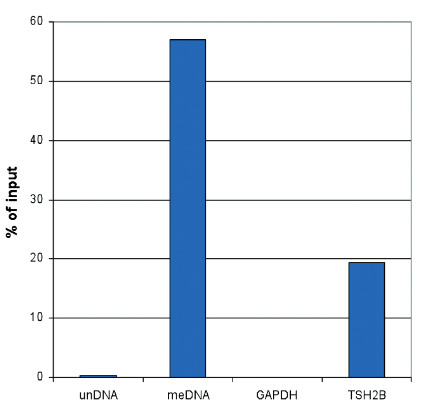
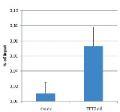
MeDIP (Methylated DNA immunoprecipitation) was performed on 1 μg fragmented human genomic DNA using 0.2 μg of the monoclonal antibody against 5-mC and the MagMeDIP Kit. The fragmented DNA was spiked with the internal controls present in the kit (methylated DNA (meDNA) as a positive and unmethylated DNA (unDNA) as a negative control) prior to performing the IP. QPCR was performed with optimized primer sets, included in the kit, specific for the methylated and unmethylated DNA controls, and for a known methylated (TSH2B) and unmethylated (GAPDH) genomic region. The recovery is expressed as a % of input (the relative amount of immunoprecipitated DNA compared to input DNA after qPCR analysis).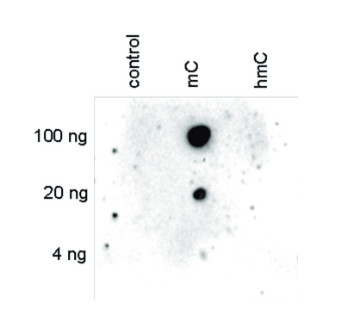
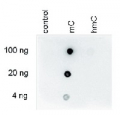
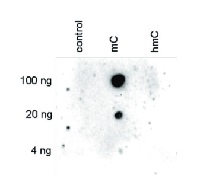
Dot blot analysis was performed using the hmC, mC and C controls. One hundred to 4 ng (equivalent of 5 to 0.2 pmol of C-bases) of the controls were spotted on a membrane. The antibody was used at a dilution of 1:250. This Dot blot shows a high specificity of the antibody for the methylated control.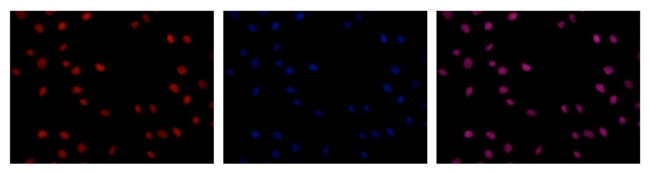
Immunofluorescence: HeLa cells were stained with the monoclonal antibody against 5-mC and with DAPI. Cells were fixed with 2.5% formaldehyde for 30’ and blocked with PBS/TX-100 containing 5% normal goat serum and 1% BSA. The cells were immunofluorescently labelled with the 5-mC antibody (left) diluted 1:100 in blocking solution followed by an anti-mouse antibody conjugated to Alexa594. The middle panel shows staining of the nuclei with DAPI. A merge of the two stainings is shown on the right.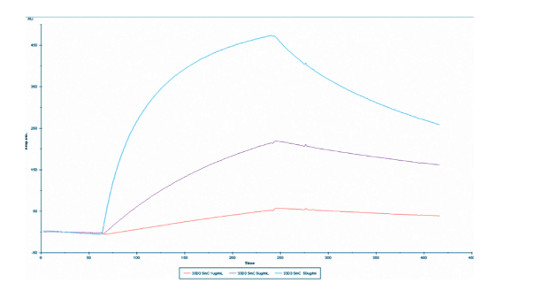
Surface plasmon resonance (SPR): A synthesized biotin-labeled 5-mC conjugate was immobilized on a CM4 BIAcore sensorchip (GE Healthcare, France). Briefly, two flowcells were prepared by sequential injections of EDC/NHS, streptavidin, and ethanolamine. One of these flowcells served as negative control (biotinylated spacer without 5-mC), while biotinylated 5-mC conjugate was injected in the other one, to get an immobilization level of 55 response units (RU). All SPR experiments were performed, using HBS-N buffer (10 mM HEPES,150 mM NaCl, pH 7.4), at a flow rate of 5 μl/min. Interaction assays involved injections of 3 different dilutions of the 5-mC monoclonal antibody over the biotinylated 5-mC conjugate and negative control surfaces, followed by a 3 min washing step with HBS-N buffer to allow dissociation of the complexes formed. At the end of each cycle, the streptavidin surface was regenerated by injection of 0.1M citric acid (pH=3). The sensorgrams correspond to the biotinylated 5-mC conjugate surface signal subtracted with the negative control. Data from the sensorgrams that reached binding equilibrium were used for Scatchard analysis. The value of the dissociation constant (kd) obtained by global fitting and 1:1 Langmuir model is 15 nM. - Additional Information
-
Additional information: This antibody is purified by gel filtration and is present in PBS containing 0.05 % sodium azide.
This antibody is very suitable for MeDIP/MeDIP-seq (methylated DNA immunoprecipitation) applications. - Background
-
Background: 5-mC (5-methylcystosine) is a methylated form of the DNA base cytosine. Methylation may be involved in the regulation of gene transcription.
Alternative names: 5 Me citidine, 5 Methycytosine, 5 Me Cytidine, methyl CpG. - Reviews:
-
This product doesn't have any reviews.