1
AtpH | ATP synthase subunit c (chloroplastic)
AS05 071 | Clonality: Polyclonal | Host: Rabbit | Reactivity: A. thaliana, S. oleracea, N. benthamina, T. elongatus
- Product Info
-
Immunogen: Purified c subunit from Spinacia oleracea UniProt: P69447.
Host: Rabbit Clonality: Polyclonal Purity: Serum Format: Lyophilized Quantity: 200 �l Reconstitution: For reconstitution add 200 �l of sterile water Storage: Store lyophilized/reconstituted at -20�C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Western blot (WB) Recommended dilution: 1 : 1000-1 : 10 000 (WB) Expected | apparent MW: 8 kDa (for Arabidopsis thaliana)
- Reactivity
-
Confirmed reactivity: Arabidopsis thaliana, Nicotiana benthamina, Spinacia oleracea, Thermosynechococcus elongatus Predicted reactivity: Algae, Cannabis sativa, Glycine max, Oryza sativa, Physcomitrium patens, Pisum sativum, Populus alba, Pinus thunbergii, Zea mays, Vitis vinifera
Species of your interest not listed? Contact usNot reactive in: No confirmed exceptions from predicted reactivity are currently known - Application Examples
-
Application example
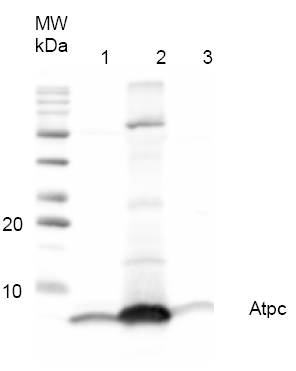
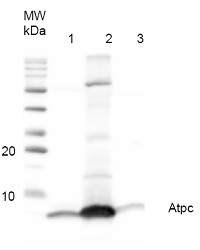
0.7 μg of purified ATP-synthase complex from Spinacia oleracea (CF0F1) (1), 15.3 µg of purified ATP-synthase complex (CF0F1) from Nicotiana benthamiana (2) and 48.6 µg of Thermosynechococcus elongatus thylakoid preparation (3) were separated on 12% polyacrylamide gel and blotted on PVDF membrane. Filters where blocked (0.5h), incubated with 1: 1000 anti-AtpH antibodies (1h), followed by incubation with 1: 5 000 secondary anti-rabbit antibody (1.25h), coupled to HRP and visualized with chemiluminescence detection reagent.
- Additional Information
-
Additional information: This product can be sold containing proclin if requested.
Additional information (application): Note that increased incubation at 95�C (20-30 min) prior to loading is recommended to break the multimeric c-mer structure, detection of partial ring structures (e,g, 5 or 6 subunits) may occur - Background
-
Background: F-type ATPase (ATP synthase) is the universal enzyme that synthesizes ATP from ADP and phosphate using the energy stored in a transmembrane ion gradient. Multiple copies of the c subunit build up the ring structure (in spinach a 14-mer of ~112 kDa) of the membrane bound Fo-part of the enzyme.
- Product Citations
-
Selected references: Nath et al. (2016). A Nitrogen-Fixing Subunit Essential for Accumulating 4Fe-4S-Containing Photosystem I Core Proteins. Plant Physiol. 2016 Dec;172(4):2459-2470. Epub 2016 Oct 26.
Lawrence et al. (2010). Recombinant production and purification of the subunit c of chloroplast ATP synthase. Protein Expression and Purification 76: 15-24.
- Protocols
-
Agrisera Western Blot protocol and video tutorials
Protocols to work with plant and algal protein extracts - Reviews:
-
This product doesn't have any reviews.