1
PsbA | D1 protein of PSII, DE-loop
AS10 704 | Clonality: Polyclonal | Host: Rabbit | Reactivity: A.thaliana, C.reinhardii, H.vulgare, M.truncatula, N. oceanica, N. oleoabundans UTEX 1185 (chlorophyta), P.patens, P.sativum, S.vulgaris, S.alba, S.oleracea, Synechococcus sp. PCC 7942, Synechocystis sp. PCC6803, Triticum sp.
- Product Info
-
Immunogen: KLH-conjugated synthetic peptide, amino acids 234-242 of Arabidopsis thaliana D1 protein UniProt: P83755, TAIR:AtCg00020
Host: Rabbit Clonality: Polyclonal Purity: Serum Format: Lyophilized Quantity: 50 µl Reconstitution: For reconstitution add 50 µl of sterile water Storage: Store lyophilized/reconstituted at -20°C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Blue Native PAGE (BN-PAGE), Western blot (WB) Recommended dilution: 1 : 10 000, thylakoid fraction (WB) Expected | apparent MW: 38 | 28-30 kDa
- Reactivity
-
Confirmed reactivity: Arabidopsis thaliana, Chlamydomonas reinhardii, Hordeum vulgare, Medicago truncatula, Nannochloropsis oceanica strain IMET1, Neochloris oleoabundans UTEX 1185 (chlorophyta), Nicotiana tabacum, Physcomitrium patens, Pisum sativum, Silene vulgaris, Sinapsis alba, Spinacia oleracea, Synechococcus sp. PCC 7942, Synechocystis sp. PCC6803, Triticum sp. Predicted reactivity: Cucumis sativus, Glycine max, Nannochloropsis sp., Oryza sativa, Populus balsamifera, Ricinus communis, Zea mays, Vitis vinifera
Species of your interest not listed? Contact usNot reactive in: No confirmed exceptions from predicted reactivity are currently known - Application Examples
-
Application example
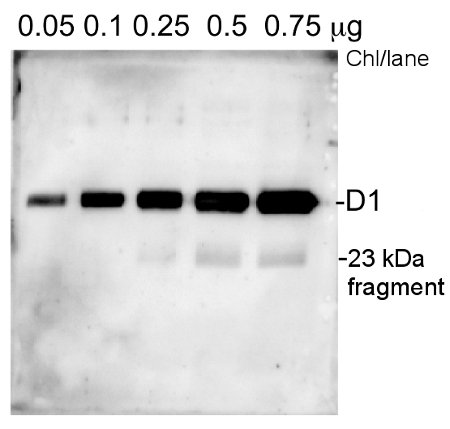
Thylakoid membranes from Arabidopsis (0.05-0.75 µg of Chl) were separated on 14%AA+ 6M urea gels and blotted 1h to PVDF. Blots were blocked immediately following transfer in 5% milk solution for 1h at room temperature with agitation. Blots were incubated in the primary antibody at a dilution of 1: 20 000 o.n. at 4°C with agitation. The antibody solution was decanted and the blot was rinsed briefly twice, then washed once for 15 min and 3 times for 5 min in TBS-T at room temperature with agitation. Blots were incubated in secondary antibody, HRP conjugated, diluted to 1:5 000. The blots were washed as above and developed for 5 min with ECL-Plus detection reagent according to the manufacturers instructions. Exposure time was 1 min in CCD camera Fuji4000.
Courtesy Professor Cornelia Spetea-Wiklund, University of Ghotenburg, Sweden
Application examples: 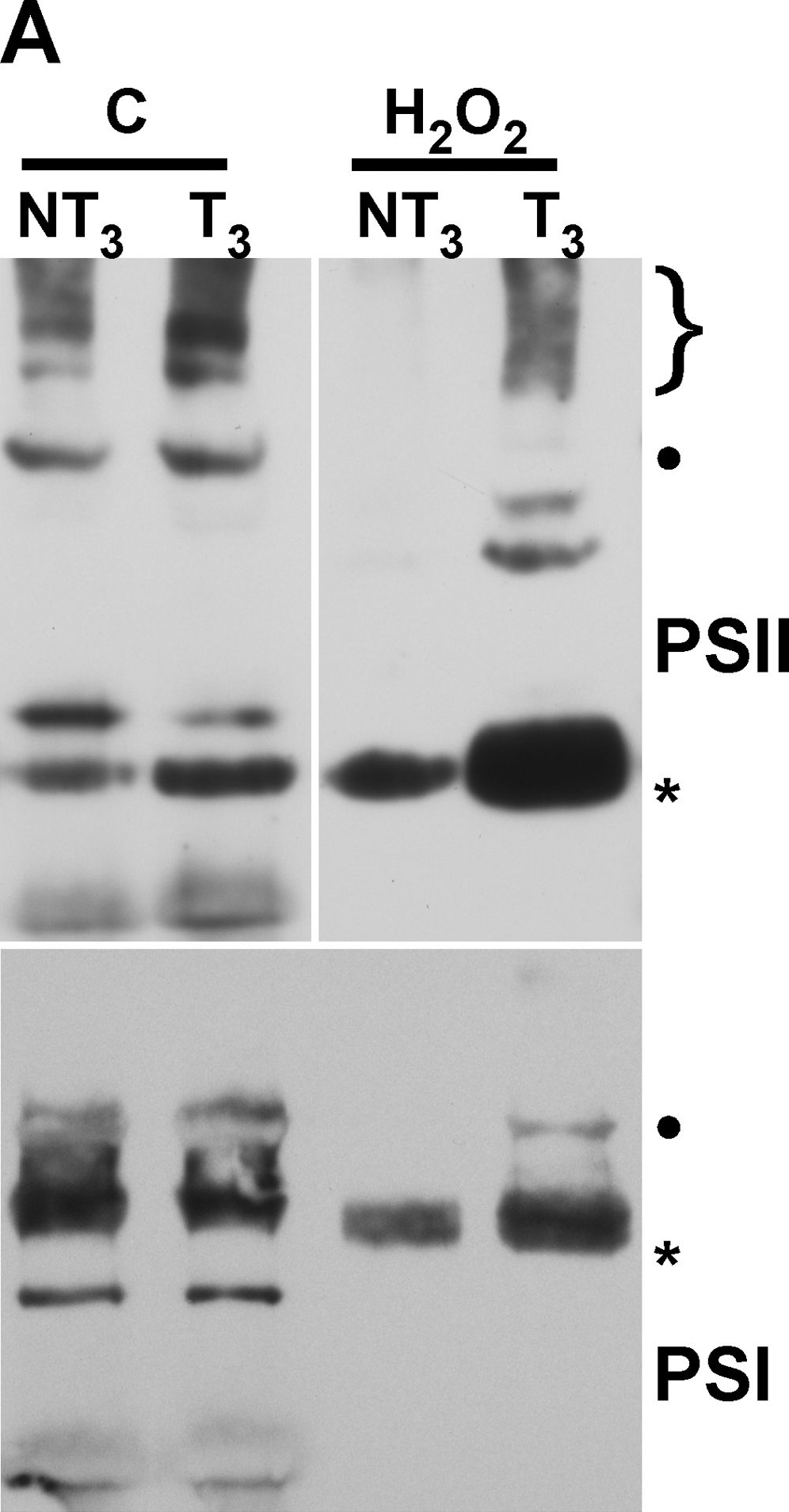
Reactant: Nicotiana tabacum (Common tobacco)
Application: Western Blotting
Pudmed ID: 23227265
Journal: PLoS One
Figure Number: 4A
Published Date: 2012-12-12
First Author: Almoguera, C., Prieto-Dapena, P., et al.
Impact Factor: 2.942
Open PublicationMembrane protein complexes of PSII and PSI survive in the 35S:A9 seedlings.(A) Western detection of complexes after the H2O2 treatments. The PSII complexes (top) separated by BN-PAGE were detected using anti-D1 (DE-loop) antibodies at 1/5,000 dilution. The PSI complexes (bottom) were detected using anti-PsaB antibodies at 1/4,000 dilution. PSII symbols: •, and the bracket on top respectively mark the dimeric PSII complex, and the PSII-LHCII super-complexes. The asterisk marks the CVII (CP43-less PSII monomer) complex. PSI symbols: • marks the PSI-LHCI super-complex that co-migrates in our gel system with the dimeric PSII complex; the asterisk marks the PSI monomer. (B) The PSII complexes also withstand drastic dehydration. The thylakoid samples were analyzed immediately (0 h) after the dehydration treatment (DT), and following rehydration for 16 h, DT (16 h). In this case the complexes were detected using anti-D1 (C-terminal) antibodies at 1/15,000 dilution. An additional PSII complex mentioned in the text is indicated: CV (**, PSII monomer).
Reactant: Arabidopsis thaliana (Thale cress)
Application: Western Blotting
Pudmed ID: 25835989
Journal: PLoS One
Figure Number: 3B
Published Date: 2015-04-04
First Author: Fristedt, R., Martins, N. F., et al.
Impact Factor: 2.942
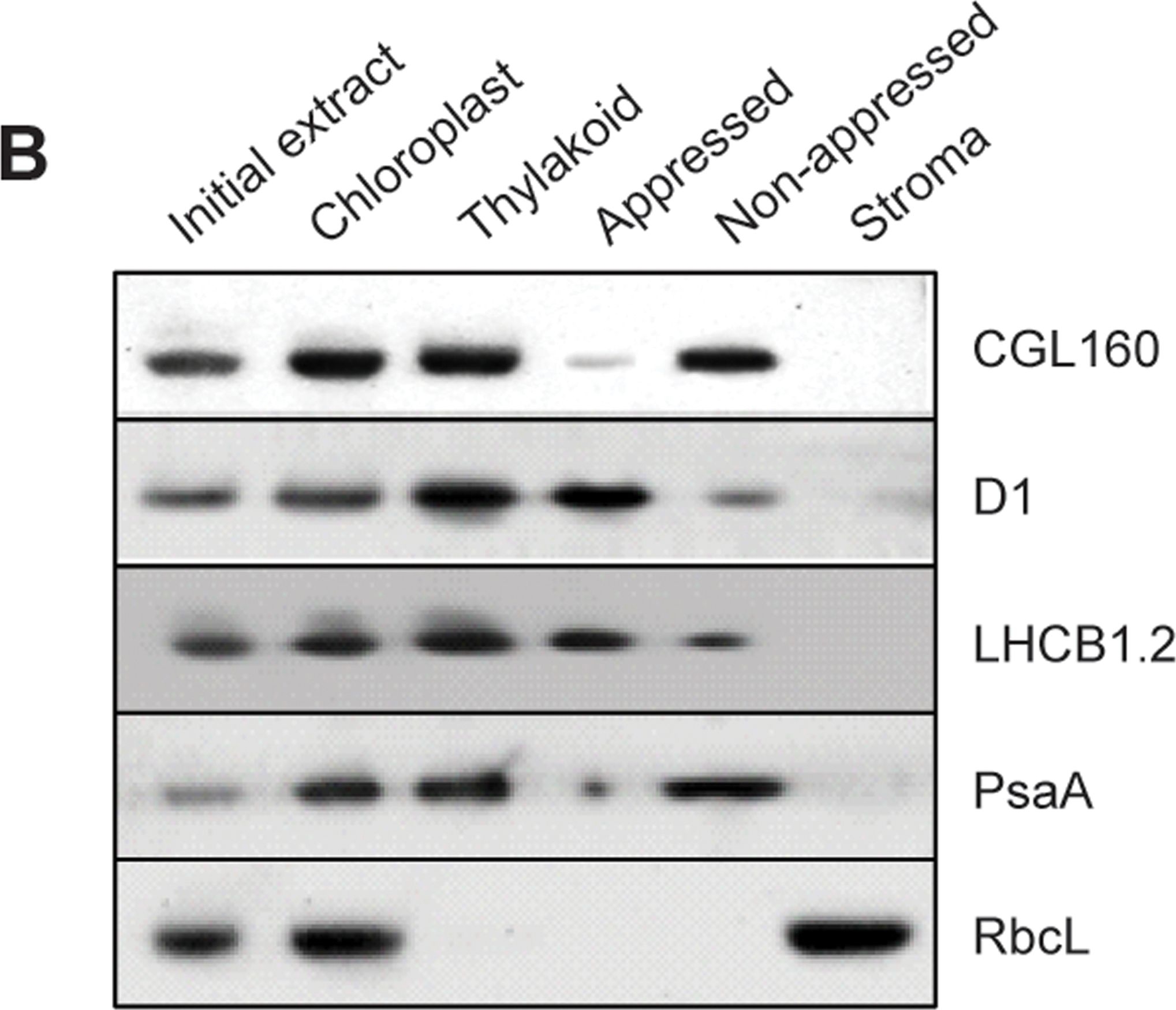
Open PublicationCGL160 is a membrane-integral chloroplast protein located in non-appressed thylakoid membranes.A. Subcellular localization of CGL160. Immunoblot analysis performed on initial plant extract, isolated chloroplasts or isolated mitochondria. TOM40 is a marker for the mitochondrion. B. Suborganellar localization determined by immunoblot analysis of chloroplast proteins diagnostic for photosystem I (PsaA), photosystem II (D1), Rubisco, LHCII (LHCB1.2) and CGL160. Protein extracts were separated by SDS-PAGE and probed with specific antibodies directed against PsaA, (PSI reaction center subunit), D1 (PSII reaction center subunits), LHCB1 (outer PSII antenna protein), the large subunit of Rubisco, a soluble protein in the stroma, and CGL160. C. Immunoblot analysis show that CG160 is an integral membrane protein associated with the thylakoid membranes. Isolated thylakoid membranes were washed with 0.4 M NaCl, and the thylakoid membranes and the supernatant were probed by immunoblotting with antibodies against CGL160 and the PSII reaction center protein D1.
Reactant: Plant
Application: Western Blotting
Pudmed ID: 27590049
Journal: BMC Plant Biol
Figure Number: 9A
Published Date: 2016-09-02
First Author: Mazur, R., Sadowska, M., et al.
Impact Factor: 4.142
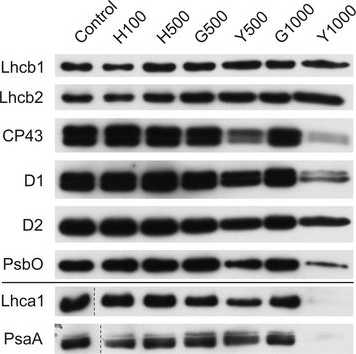
Open PublicationChanges of PSII and PSI antenna and core protein levels. Proteins from control and Tl-treated white mustard leaves were separated by SDS-PAGE followed by immunodetection with antibodies against Lhcb1, Lhcb2, Lhca1 (antenna proteins) and D1, D2, CP43, PsbO, PsaA (core proteins). Samples were loaded on the equal amount of chlorophyll (0.25 ?g). Description of samples abbreviation as given in the legend to Fig. 3
- Additional Information
-
Additional information: Antibody will detect 23 kDa N-terminal fragment Additional information (application): Antibody is recognizing a 23 kDa fragment in spinach and Arabidopsis thylakoidsfor usage on total cell extracts the dilution needs to be determined experimentally - Background
-
Background: The psbA gene has been cloned from many species of plants, green algae, and cyanobacteria. The psbA gene is located in the chloroplast genome and encodes for the D1 protein, a core component of Photosystem II. PsbA/D1 is rapidly cycled under illumination in all oxygenic photobionts. Tracking PsbA pools using the Global PsbA antibody can show the functional content of Photosystem II in a wide range of samples. Alternative names: 32 kDa thylakoid membrane protein, photosystem II protein D1
- Product Citations
-
Selected references: Lu et al. (2021). Role of an ancient light-harvesting protein of PSI in light absorption and photoprotection. Nat Commun. 2021 Jan 29;12(1):679. doi: 10.1038/s41467-021-20967-1. PMID: 33514722; PMCID: PMC7846763. (blue-native PAGE)
Mazur et al. (2021) The SnRK2.10 kinase mitigates the adverse effects of salinity by protecting photosynthetic machinery. Plant Physiol. 2021 Dec 4;187(4):2785-2802. doi: 10.1093/plphys/kiab438. PMID: 34632500; PMCID: PMC8644180.
Rantala et al. (2020). PGR5 and NDH-1 systems do not function as protective electron acceptors but mitigate the consequences of PSI inhibition. Biochim Biophys Acta Bioenerg. 2020 Jan 11;1861(3):148154. doi: 10.1016/j.bbabio.2020.148154.
Grieco et al. (2020). Adjustment of photosynthetic activity to drought and fluctuating light in wheat. Plant Cell Environ. 2020 Mar 16. doi: 10.1111/pce.13756.
Rantala and Tikkanen et al. (2018). Phosphorylation?induced lateral rearrangements of thylakoid protein complexes upon light acclimation. Plant Direct Vol. 2, Issue 2. - Protocols
-
Agrisera Western Blot protocol and video tutorials
Protocols to work with plant and algal protein extracts
Agrisera Educational Posters Collection
- Reviews:
-
Martina Giovanardi | 2016-10-19We used this antibody on the green microalga Neochloris oleoabundans and it worked really well. Loading 0.5 �g of Chlorophyll we got very strong signal without aspecific bands.Jun Liu | 2014-01-14This antibody has been working well. It always gave a strong signal around 30 kDa in Arabidopsis, wheat and rice.
Accessories
AS01 016 | Clonality: Polyclonal | Host: Hen | Reactivity: [global antibody] for higher plants, algae, cyanobacteria and dinoflagellates
Benefits of using this antibody
AS05 084 | Clonality: Polyclonal | Host: Rabbit | Reactivity: [global antibody] for higher plants, algae, liverwort, cyanobacteria, diatoms | cellular [compartment marker] of thylakoid membrane
Benefits of using this antibody

