1
cFBPase | Cytosolic fructose-1,6-bisphosphatase (cytoplasm marker in photosynthetic tissues)
AS04 043 | Clonality: Polyclonal | Host: Rabbit | Reactivity: Arabidopsis thaliana, Brassica napus, Macroptilium atropurpureum, Nicotiana benthamiana, Pinus silvestris, Pinus yunanniensis, Oryza sativa, Petunia hybrida cv. Mitchell, Solanum tuberosum, Zea mays | cellular [compartment marker] of cytoplasm
- Product Info
-
Immunogen: Overexpressed cytosolic fructose 1,6 bisphosphatase (cFBPase) derived from the sequence from Arabidopsis thaliana cFBPase UniProt: Q9MA79, TAIR: AT1G43670
Host: Rabbit Clonality: Polyclonal Purity: Serum Format: Lyophilized Quantity: 50 µl Reconstitution: For reconstitution add 50 µl of sterile water Storage: Store lyophilized/reconstituted at -20°C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Immunolocalization (IL), Western blot (WB) Recommended dilution: 1: 500 (IL), 1 : 5 000 (WB) Expected | apparent MW: 45 | 37 kDa (Arabidopsis thaliana)
- Reactivity
-
Confirmed reactivity: Arabidopsis thaliana, Brassica napus, Macroptilium atropurpureum, Nicotiana benthamiana, Pinus silvestris, Pinus yunanniensis, Oryza sativa, Petunia hybrida cv. Mitchell, Solanum tuberosum, Zea mays
Predicted reactivity: Capsella rubella, Pisum sativum, Ricinus communis, Glycine max, Phaseolus vulgaris, Sesamum indicum, Spinacia oleracea, Populus trichocarpa, Vitis vinifera
Species of your interest not listed? Contact usNot reactive in: Chlamydomonas reinhardtii
- Application Examples
-
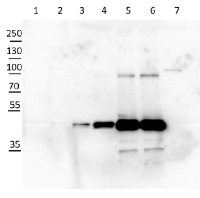
Application example 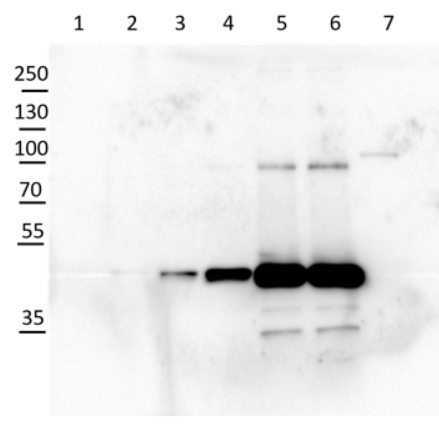
10 µg of Arabidopsis thaliana Col-0 WT chloroplast total protein (1), 10µg Col-0 WT chloroplast stroma protein (2), Col-0 WT total leaf sample 1:10 dilution (3), Col-0 WT total leaf sample 1:2 dilution (4), Col-0 WT total leaf sample undiluted (5), Col-0 WT total leaf sample undiluted (6) and recombinat plastidial FBPase 0.05 µg, expressed in E.coli with no cTP present in the sequence (7), extracted with 2x Laemmli buffer and denatured at 95°C for 5 min. were separated on 10% SDS-PAGE and blotted to Millipore Immobilon-P membrain (carried out at 100 V for 90 min at 4°C in blotting buffer (25 mM Tris-HCl, 192 mM glycine, 10 % [v/v] methanol). Blots were blocked with blocking solution ( TBST buffer (20 mM Tris-HCl pH 7.4, 150 mM NaCl, 0.05 % [v/v] Tween-20) supplemented with 5 % [w/v] milk powder) for 1h at room temperature (RT) with agitation. Blot was incubated in the primary antibody at a dilution of 1: 5 000 for 15h (over night) at 4°C with agitation in TBST. The antibody solution was decanted and the blot was washed 6 times for 10 min in TBS-T at RT with agitation. Blot was incubated in secondary antibody (anti-rabbit IgG horse radish peroxidase conjugated) diluted to 1:20 000 in for 1h at RT with agitation. The blot was washed as above and ChemiGlow West Chemiluminescence substrate was used for development according to the manufacturer’s instructions and imaged using the ChemiDoc imaging system (Biorad, Cressier, France). Exposure time was 15 seconds.
Courtesy of Zanella Martina, ETH Zürich, Switzerland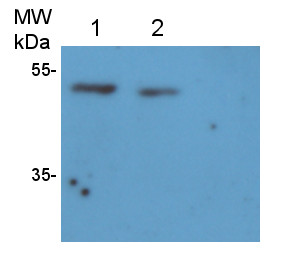
2 µg of total protein from Arabidopsis thaliana roots crude extract (1), supernatant after 10 0000 g centrifugation (2), extracted with ice-cold extraction buffer [50 mM Tris-HCl pH 7.5, 0.33 M Sucrose, 5 mM EDTA, 1x proteinase inhibitor] and denatured gradually with [50 mM Tris (pH 6.8), 10% Glycerol, 2% SDS, 2.5 M Urea, 0.005% Bromophenol Blue] at first with 55°C for 15 min followed by 95°C for 5 min. Proteins were separated on 12 % SDS-PAGE. Proteins were blotted 1h to PVDF using semi-dry transfer. Blots were blocked with 5 % nonfat milk + 0.1 % BSA for 1h at room temperature (RT) with agitation. Blot was incubated in the primary antibody at a dilution of 1: 35 000 for ON/4°C with agitation in TBS-T. The antibody solution was decanted and the blot was rinsed briefly twice, then washed once for 15 min and 3 times for 5 min in TBS-T at RT with agitation. Blot was incubated in secondary antibody (anti-rabbit IgG horse radish peroxidase) diluted to 1:1 000 in for 2h at RT with agitation. The blot was washed as above and developed for 5 min withchemiluminescent detection reagent. Exposure time was 20 min.
Courtesy of Dr. Joanna Jeleńska and Dequantarius Speed, University of Chicago, USA
Application examples: 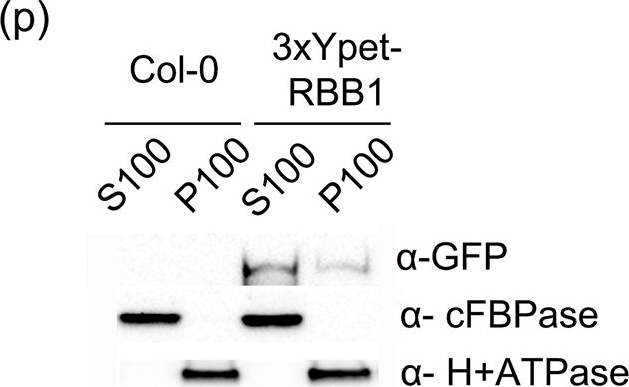
Reactant: Arabidopsis thaliana (Thale cress)
Application: Western Blotting
Pudmed ID: 25915922
Journal: PLoS One
Figure Number: 6p
Published Date: 2015-04-29
First Author: Han, S. W., Alonso, J. M., et al.
Impact Factor: 2.942
Open Publication3xYpet-RBB1 is a cytoplasmic protein that associates with the tonoplast.(a-c) A 3xYpet-RBB1 fusion complements the bulb phenotype of rbb1-2. Four-day-old seedlings from rbb1-2 (a) or rbb1-2 3xYpet-RBB1 (b, c) were stained with Lysotracker Red (magenta, a, b) to label the vacuole. The signal from 3xYpet-RBB1 (c) for the complemented plant is shown. (d-f) 3xYpet-RBB1 does not co-localize with FM4-64. Images show the cotyledon of 7-d-old seedlings expressing 3xYpet-RBB1 (d) stained with 5 ?M FM4-64 (e) and the merged image (f). (g-i) 3xYpet-RBB1 and RFP-SYP22 do not co-localize. Cotyledons of 7-d-old seedlings expressing 3xYpet-RBB1 (g) and RFP-SYP22 (h) were imaged. Merged image (i) is shown. (j-l) 3xYpet-RBB1 and Lysotracker Red do not co-localize in rosette leaves. Seedlings from the 3xYpet-RBB1 line were stained with Lysotracker Red. Signal from 3xYpet-RBB1 (j), Lysotracker Red (k) and the merged image (l) are shown. Arrowheads indicate the localization of 3xYpet-RBB1 at the tip of elongating trans-vacuolar strands. (m-o) GFP molecules can also label the tips of transvacuolar strands. Seedlings from a 35::GFP marker line were stained with Lysotracker Red. The GFP signal (m), Lysotracker Red (n) and the merged image (o) are shown. Arrowheads indicate the tip of TVS. All scale bars = 20 ?m. (p) 3xYpet-RBB1 accumulates in the soluble (S100) and membrane pellet (P100) fractions from whole seedlings. Immunoblot of 3xYpet-RBB1 soluble (S100) and membrane (P100) fractions from Col-0 and 3xYpet-RBB1 transgenic plants using antibodies against GFP (?-GFP), cFBPase (?-cFBPase, control for soluble fraction), and Plasma Membrane H+ATPase (?-H+ATPase, control for membrane fraction).
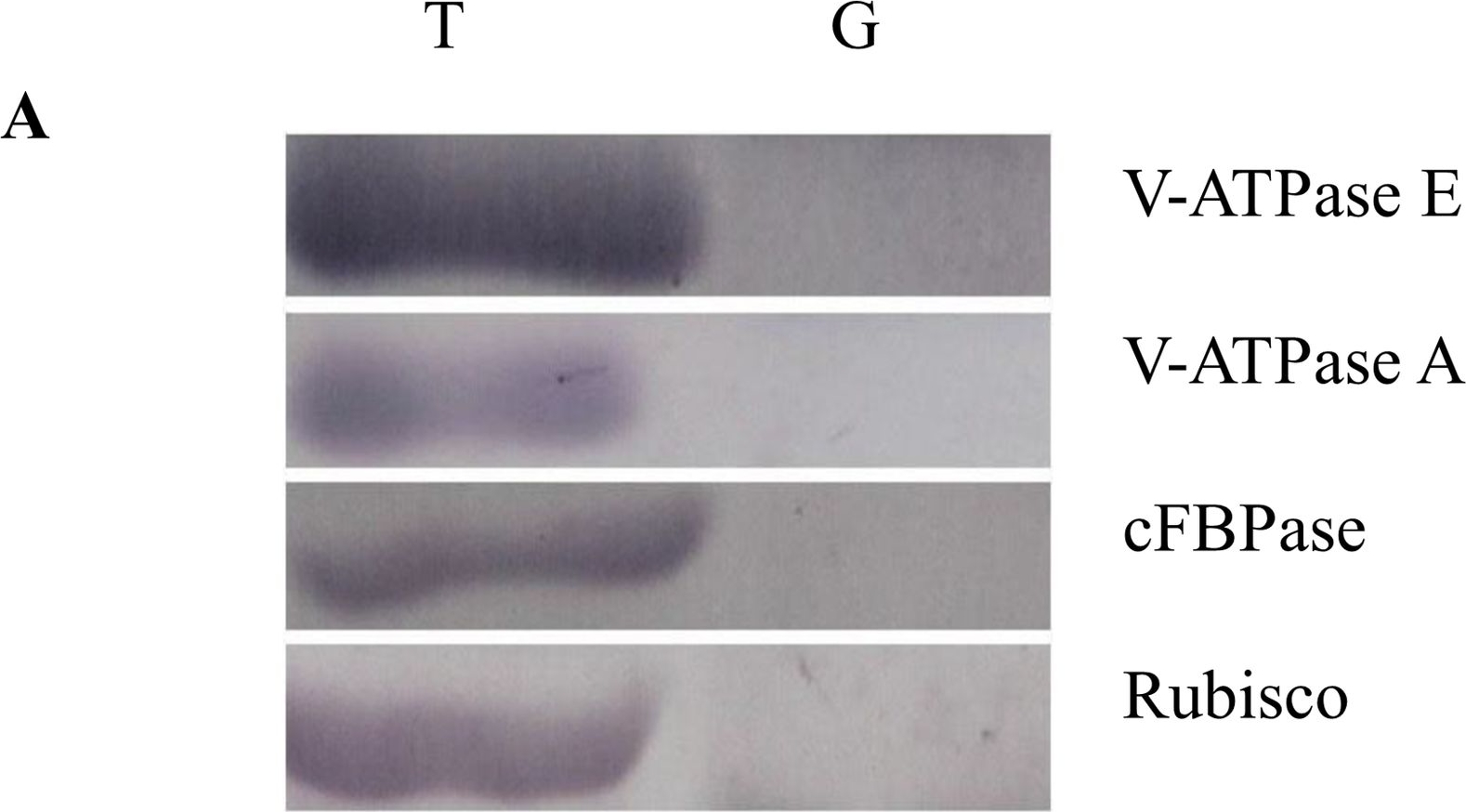
Reactant: Oryza sativa (Asian rice)
Application: Western Blotting
Pudmed ID: 27992503
Journal: PLoS One
Figure Number: 4A
Published Date: 2016-12-20
First Author: Xing, S., Meng, X., et al.
Impact Factor: 2.942
Open PublicationWestern blot image of the starch granule proteome. Same amount of proteins (25 ?g per lane) were loaded.A: Western blot images with different antibodies. The anti-bodies used were for V-ATPase E, V-ATPase A, Anti-Rubisco, cFBPase (Agrisera, Sweden). T: Total proteins; G: Starch granule proteins. B: Western blot image of protein acetylation of endosperm and starch granule proteins. Antibodies for acetylated Lysine (ImmuneChem) were used for Western blots. The source of proteins is indicated on the top of the lane. M: Protein marker; T: Total proteins extracted from endosperm; G: Proteins extracted from starch granules.
- Additional Information
-
Additional information: Kinetic and allosteric properties of the plant cytosolic FBPase are remarkably similar to the mammalian and yeast FBPase, but differ greatly from those of the chloroplastic FBPase. The antibody could detect FBPase from the human COS-7 cell line transfected with FBP1 expressing vector.
This product can be sold containing ProClin if requested.Additional information (application): This antibody does not react with chloroplastic form of FBPase.
Will this antibody be good as a cytosolic (non-microsomal control) in Arabidopsis thaliana roots? Although it has never been tested there is every likelihood that cFBPase will be expressed at reasonable levels even in roots. Even though the biosynthetic flux through to Sucrose may not be high as in mesophyll cells, central metabolism will still be active in young roots and the Sucrose etc being supplied externally still needs to be utilised. - Background
-
Background: Fructose-1,6 bisphosphatase (FBPase) (EC=3.1.311) is one of the regulatory enzymes in the sucrose biosynthetic pathway. In non-photosynthetic tissues, it regulates the rate of gluconeogenesis. In photosynthetic tissues, two FBPase isozymes (chloroplastic and cytosolic) play key roles in carbon assimilation and metabolism. In photosynthetic tissues cFBPase (cytosolic fructose 1,6 bisphosphatase) converts triose phosphates from the chloroplast to sucrose during light hours. Alternative name: D-fructose-1,6-bisphosphate 1-phosphohydrolase
- Product Citations
-
Selected references: Singh, Muthamilarasan, Prasad (2022). SiHSFA2e regulated expression of SisHSP21.9 maintains chloroplast proteome integrity under high temperature stress. Cell Mol Life Sci. 2022;79(11):580. Published 2022 Nov 3. doi:10.1007/s00018-022-04611-12
Cui, Liu, Li, et al. (2022) The cellulose--lignin balance affects the twisted growth of Yunnan pine trunk. Authorea. October 10, 2022. DOI: 10.22541/au.166538021.18232197/v2
Wang et al. (2022), Arabidopsis Ubiquitin-Conjugating Enzymes UBC4, UBC5, and UBC6 Have Major Functions in Sugar Metabolism and Leaf Senescence, Int. J. Mol. Sci. 2022, 23(19), 11143; https://doi.org/10.3390/ijms231911143
He, Gao, Luo, et al. (2022) VAMP724 and VAMP726 are involved in autophagosome formation in Arabidopsis thaliana [published online ahead of print, 2022 Oct 13]. Autophagy. 2022;1-18. doi:10.1080/15548627.2022.2127240
Lim et al (2022). Arabidopsis guard cell chloroplasts import cytosolic ATP for starch turnover and stomatal opening. Nat Commun. 2022 Feb 3;13(1):652. doi: 10.1038/s41467-022-28263-2. PMID: 35115512; PMCID: PMC8814037.
- Protocols
-
Agrisera Western Blot protocol and video tutorials
Protocols to work with plant and algal protein extracts - Reviews:
-
LI BAIYING | 2020-02-05This cFBAPase antibody is a very good one for western blot and shows specific and strong signals as a cytoplasm marker for Arabidopsis.Xiangfeng Wang | 2012-06-07This antibody shows very specific and strong signals for western blot using Arabidopsis materials.| 2011-12-07This antiserum is very specific and has a high titre.
Accessories

AS05 086 | Clonality: Polyclonal | Host: Rabbit | Reactivity: A. thaliana, C. annuum, C. sativus, F. margarita Swingle, F. arundinacea, H. vulgare, L. esculentum, L. chilense, Malus x domestica Borkh. c.v. Fuji, M. polymorpha, Medicago truncatula, N. tabacum, O. sativa, P. glauca, Populus sp., S. tuberosum, S. sogarandinum, Triticum | cellular [compartment marker] of cytoplasm
Replaced by AS14 2813


