1
Anti-Rubisco | 557 kDa hexadecamer
- Product Info
-
Immunogen: purified 557 kDa hexadecamer Rubisco protein complex from Spinacia oleracea (SIGMA-ALDRICH R-8000), UniProt: P00875 and Q43832
Host: Rabbit Clonality: Polyclonal Purity: Serum Format: Lyophilized Quantity: 50 µl Reconstitution: For reconstitution add 50 µl of sterile water Storage: Store lyophilized/reconstituted at -20°C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Immunolocalization (IL), Western blot (WB) Recommended dilution: 1 : 10 000-1 : 20 000 on 0,5-10 ug total cellular protein/lane and standard (WB), 1 : 500-1:1000 (IL) Expected | apparent MW: 53-55 | 53-55 kDa
- Reactivity
-
Confirmed reactivity: Anabaena sp. PCC 7120, Arabidopsis thaliana, Aucuba japonica, Fremyella diplosiphon, Glycine max, Hordeum vulgare, Manihot esculenta Crantz, Oryza sativa, Physcomitrium patens, Pisum sativum, Populus sp., Salsola laricifolia, Solanum tuberosum, Spinacia oleracea, Synechocystis sp. PCC 6803, Synechococcus sp. PCC7942, Zea mays Predicted reactivity: higher plants, algae, Anthoceros agrestis, Nannochloropsis sp. Not reactive in: Chlamydomonas reinhardtii (immulolocalization) - Application Examples
-
Application example
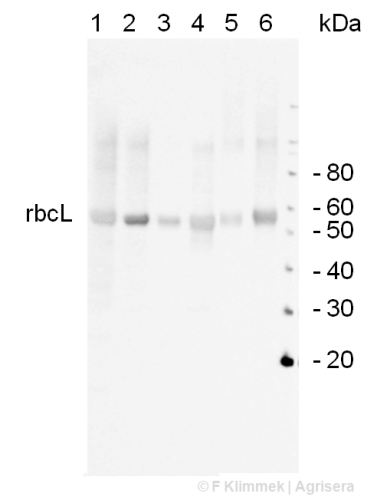
0.5 µg of total leaf protein isolated with Agrisera Protein Eextraction Buffer (AS08 300) from Arabidopsis thaliana (1), Spinacia oleracea (2), Zea mays (3), Hordeum vulgare (4), Solanum tuberosum (5), Pisum sativum (6) were separated on 4-12% NuPage (Invitrogen) LDS-PAGE and blotted 45 min (30V) to nitrocellulose. Filters where blocked 1h with 2% low-fat milk powder in TBS-T (0.1% TWEEN 20) and probed with AS07 218 (1:20 000, 1h) and secondary anti-rabbit (1:20000, 1 h) antibody (HRP conjugated) in TBS-T containing 2% low fat milk powder. Antibody incubations where followed by washings in TBS-T (15, +5, +5, +5 min). All steps were performed at RT with agitation. Signal was detected with chemiluminescence using a Fuji LAS-3000 CCD (300s, high sensitivity).
Application examples: 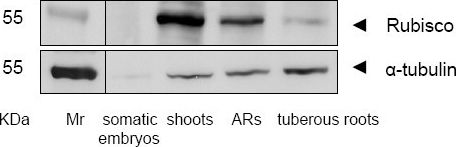
Reactant: Manihot esculenta (Cassava)
Application: Western Blotting
Pudmed ID: 20187967
Journal: Proteome Sci
Figure Number: 5A
Published Date: 2010-02-27
First Author: Li, K., Zhu, W., et al.
Impact Factor: 2.037
Open PublicationWestern blotting of Rubisco and ?-tubulan. Rubisco and ?-tubulan in the somatic embryos, plantlets and tuberous roots of cassava cultivar SC8 were detected by Western blotting using antiRubisco-polyclonal antibody from Agrisera (AS07218) and anti-?-tubulin-monoclonal antibody from Sigma-Aldrich (T9026). ARs, adventitious roots.
Reactant: Manihot esculenta (Cassava)
Application: Western Blotting
Pudmed ID: 24727655
Journal: PLoS One
Figure Number: 5A
Published Date: 2014-04-15
First Author: An, F., Fan, J., et al.
Impact Factor: 2.942
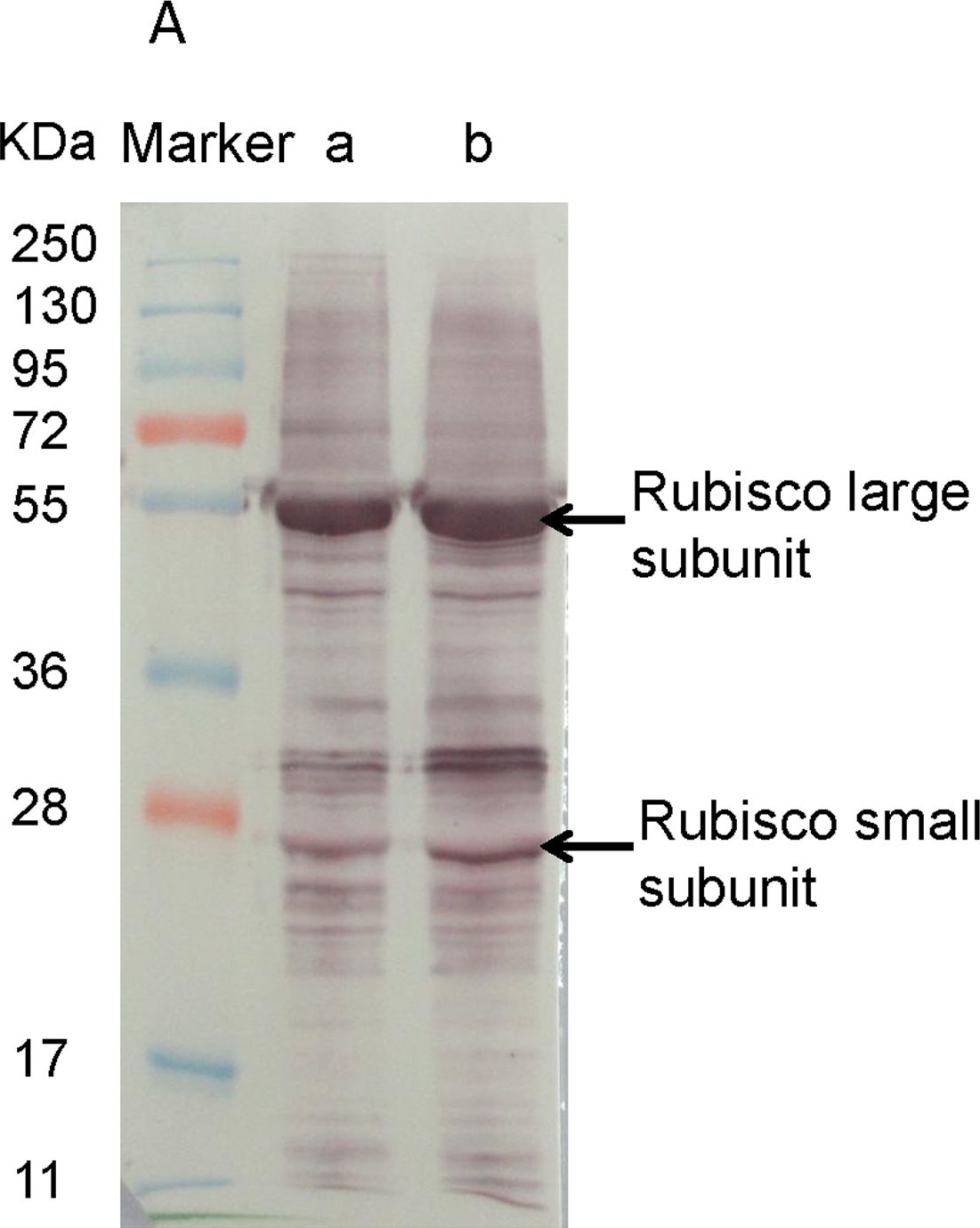
Open PublicationWestern blotting of Rubisco, APX and PrxQ.The expression of Rubisco, APX and PrxQ in leaves of cassava NZ199 diploid (a) and autotetraploid (b) genotypes were detected by western blotting using antiRubisco-polyclonal antibody (AS07218), anti-APX antibody (AS08368) and anti-PrxQ antibody (AS05093) from Agrisera, respectively.
Reactant: Manihot esculenta (Cassava)
Application: Western Blotting
Pudmed ID: 27881899
Journal: Plant Mol Biol Report
Figure Number: 8A
Published Date: 2016-11-25
First Author: An, F., Li, G., et al.
Impact Factor: 1.526
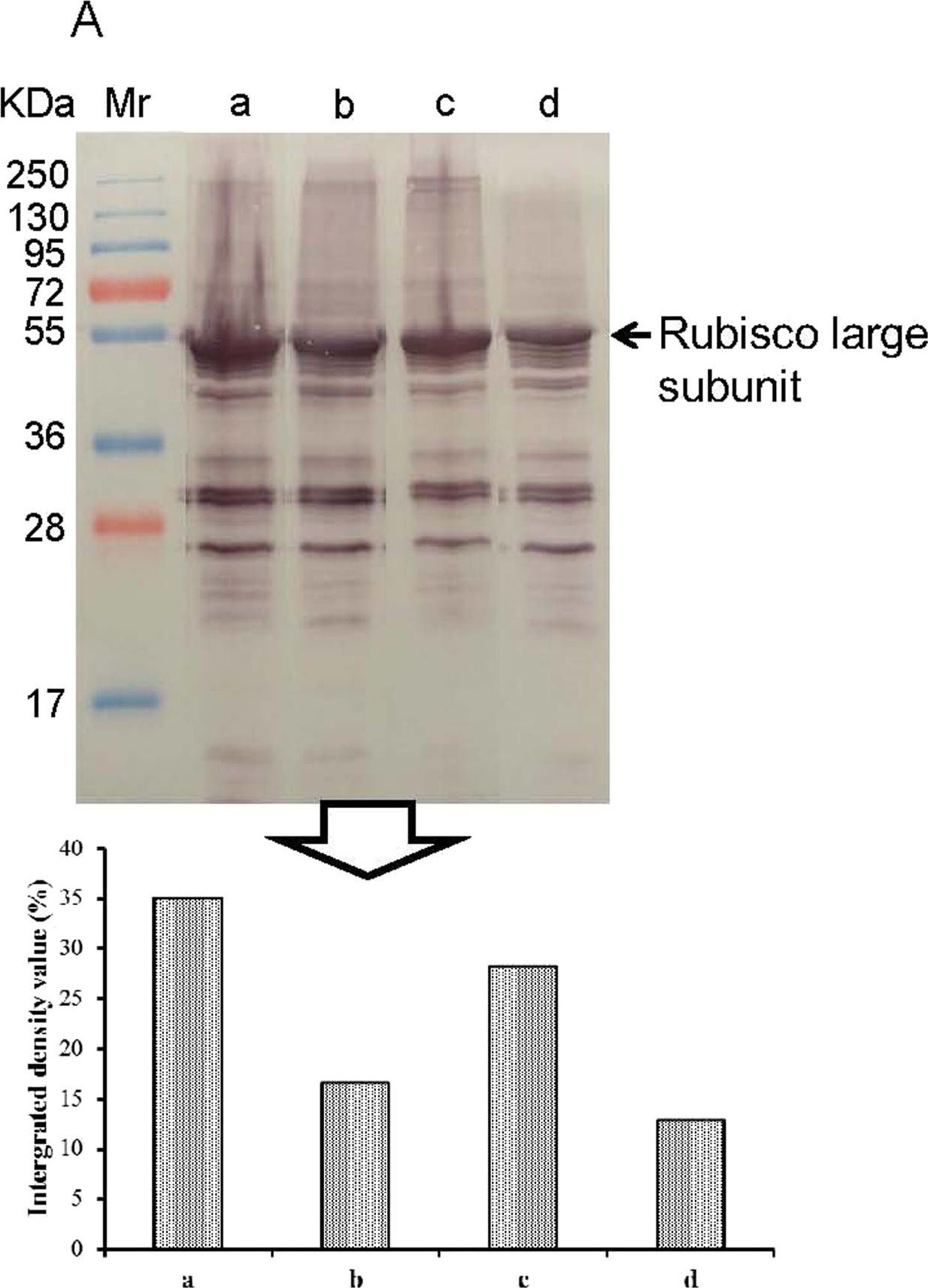
Open PublicationWestern blotting of Rubisco and peroxiredoxin. The expressions of Rubisco (a) and peroxiredoxin (b) in apical expanded leaves of SC8and Col1046 were detected by Western blotting using anti-Rubisco polyclonal antibody (AS07218) and anti-peroxiredoxin antibody (AS05093) from Agrisera, respectively. Mr, protein marker; lines a and b, the expression of Rubisco and peroxiredoxin of SC8 in 5 °C for 0 and 10 days, respectively; lines c and d, the expression of Rubisco and peroxiredoxin of Co11046 in 5 °C for 0 and 10 days, respectively
Reactant: Synechocystis
Application: Western Blotting
Pudmed ID: 31143173
Journal: Front Microbiol
Figure Number: 7A
Published Date: 2019-05-31
First Author: Oh, S. & Montgomery, B. L.
Impact Factor: 5.259
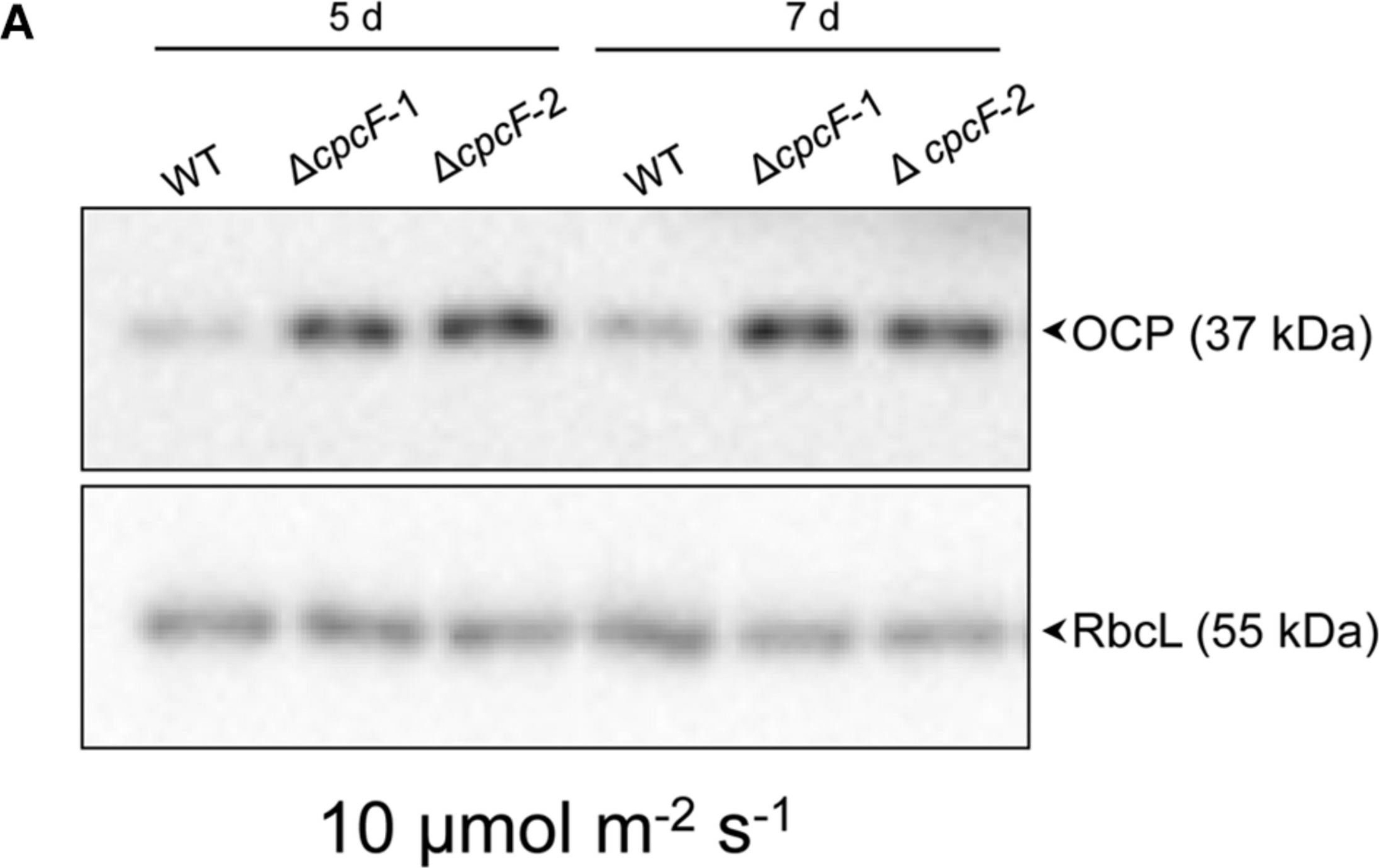
Open PublicationAccumulation of OCP protein in WT and phycobilisome-deficient strains. Immunoblot analysis was performed using anti-OCP antibody (top panels) or anti-RbcL antibody (bottom panels), with representative blots shown. Total soluble proteins were extracted from wild-type (WT), ?cpcF and ?cpcG1 strains (two independent mutant strains for each ?cpcF and ?cpcG1 included) grown under (A) 10 ?mol m–2 s–1 for 5 or 7 days or (B) 80 ?mol m–2 s–1 for 5 days of white light in BG-11/HEPES medium. 2.5 ?g of proteins were resolved on 10% (w/v) polyacrylamide gels by SDS-PAGE electrophoresis prior to immunoblotting. Arrowheads indicated OCP or RbcL proteins.
Reactant: Synechocystis
Application: Western Blotting
Pudmed ID: 31143173
Journal: Front Microbiol
Figure Number: 7B
Published Date: 2019-05-31
First Author: Oh, S. & Montgomery, B. L.
Impact Factor: 5.259
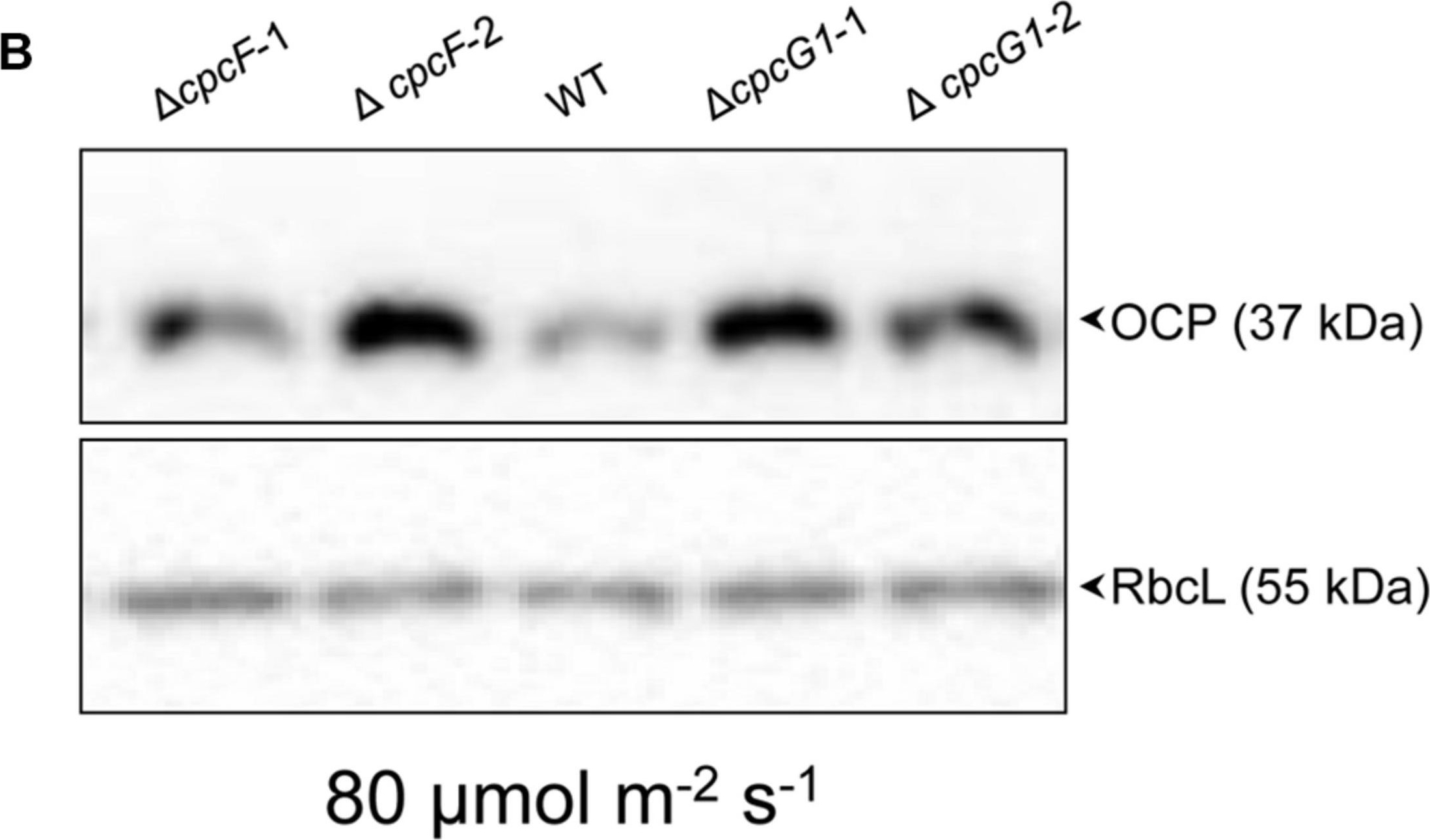
Open PublicationAccumulation of OCP protein in WT and phycobilisome-deficient strains. Immunoblot analysis was performed using anti-OCP antibody (top panels) or anti-RbcL antibody (bottom panels), with representative blots shown. Total soluble proteins were extracted from wild-type (WT), ?cpcF and ?cpcG1 strains (two independent mutant strains for each ?cpcF and ?cpcG1 included) grown under (A) 10 ?mol m–2 s–1 for 5 or 7 days or (B) 80 ?mol m–2 s–1 for 5 days of white light in BG-11/HEPES medium. 2.5 ?g of proteins were resolved on 10% (w/v) polyacrylamide gels by SDS-PAGE electrophoresis prior to immunoblotting. Arrowheads indicated OCP or RbcL proteins.
- Additional Information
-
Additional information (application): RbcS subunit is not detected by this antibody
This product can be sold containing proclin if requested - Background
-
Background: Rubisco (Ribulose-1,5-bisphosphate carboxylase/oxygenase) catalyzes the rate-limiting step of CO2 fixation in photosynthetic organisms. It is demonstrably homologous from purple bacteria to flowering plants and consists of two protein subunits, each present in 8 copies. In plants and green algae, the large subunit (~55 kDa) is coded by the chloroplast rbcL gene, and the small subunit (15 kDa) is coded by a family of nuclear rbcS genes.
- Product Citations
-
Selected references: Wang et al. (2024). Dominant-negative chaperonin mutation ptCPN60α1S57F uncovers redundancy in chloroplast rRNA processing. Plant J. 119(6):2937-2950.
Mihara et al. (2019). Thioredoxin targets are regulated in heterocysts of cyanobacterium Anabaena sp. PCC 7120 in a light-independent manner. J Exp Bot. 2019 Dec 21. pii: erz561. doi: 10.1093/jxb/erz561.
Sedaghatmehr et al. (2019). A regulatory role of autophagy for resetting the memory of heat stress in plants. Plant Cell Environ. 2019 Mar;42(3):1054-1064. doi: 10.1111/pce.13426.
Rohnke et al. (2018). RcaE-Dependent Regulation of Carboxysome Structural Proteins Has a Central Role in Environmental Determination of Carboxysome Morphology and Abundance in Fremyella diplosiphon. Mol Biol and Physiol. Vol. 3, Issue 1. DOI: 10.1128/mSphere.00617-17
Zhang et al. (2017). Composition of photosynthetic pigments and photosynthetic characteristics in green and yellow sectors of the variegated Aucuba japonica ‘Variegata’ leaves. Flora, Vol 240, March 2018, Pages 25–33.
Wang et al. (2017). Re-creation of a Key Step in the Evolutionary Switch from C3 to C4 Leaf Anatomy. Curr Biol. 2017 Nov 6;27(21):3278-3287.e6. doi: 10.1016/j.cub.2017.09.040.
Wen and Zhang (2014). Salsola laricifolia, another C 3 –C 4 intermediate species in tribe Salsoleae s.l. (Chenopodiaceae). Photosynth Res. 2014 Sep 17. (immunolocalization)
Feifei et al. (2014). Comparison of Leaf Proteomes of Cassava (Manihot esculenta Crantz) Cultivar NZ199 Diploid and Autotetraploid Genotypes. PLoS One. 2014 Apr 11;9(4):e85991. doi: 10.1371/journal.pone.0085991. eCollection 2014. - Protocols
-
Agrisera Western Blot protocol and video tutorials
Protocols to work with plant and algal protein extracts
Agrisera Educational Posters Collection
- Reviews:
-
This product doesn't have any reviews.
Related products
AS03 037 | Clonality: Polyclonal | Host: Rabbit | Reactivity: global antibody and compartment marker for higher plants, lichens, algae, cyanobacteria, dinoflagellates, diatoms
Benefits of using this antibody

