1

Anti-UGPase | UDP-glucose pyrophosphorylase (cytoplasm marker)
AS14 2813 | Clonality: Polyclonal | Host: Rabbit | Reactivity: A. thaliana, H.vulagre, Z.mays
- Product Info
-
Immunogen: His-tagged, full length Hordeum vulgare UGPase, overexpressed and purified from E.coli, UniProt: Q43772.1 Host: Rabbit Clonality: Polyclonal Purity: Serum Format: Lyophilized Quantity: 50 µl Reconstitution: For reconstitution add 50 µl of sterile water Storage: Store lyophilized/reconstituted at -20°C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Western blot (WB) Recommended dilution: 1 : 10 000 (WB) Expected | apparent MW: 52 kDa
- Reactivity
-
Confirmed reactivity: Arabidopsis thaliana, Hordeum vulgare, Zea mays
Predicted reactivity: Bambusa oldhamii, Brassica pekinensis, Brassica rapa, Capsicum annuum, Cucumis sativus, Dendrobium catenatum, Dendrocalamus sinicus, Glycine max, Gossipium hirsutum, Lycopersicum esculentum, Lycopersicum chilense, Malus domestica, Marchantia polymorpha, Oryza sativa, Picea glauca, Populus sp., Solanum tuberosum, Populus tremula, Ricinus communis, Saccharum officinarum, Vitis vinifera
Species of your interest not listed? inquire
Species of your interest not listed? Contact usNot reactive in: Homo sapiens - Application Examples
-
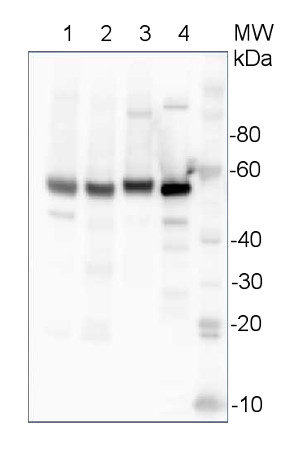
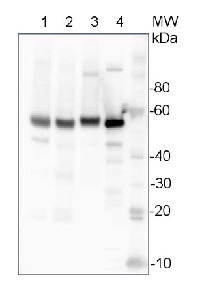
10 µg of total protein from Arabidopsis thaliana leaf (1) , Hordeum vulgare leaf (2), Zea mays leaf (3), recombinant UGPase 0.5 pmol (4), were extracted with Protein Extraction Buffer PEB (AS08 300). Samples were diluted with 1X sample buffer (NuPAGE LDS sample buffer (Invitrogen) supplemented with 50 mM DTT and heat at 70°C for 5 min and keept on ice before loading. Protein samples were separated on 4-12% Bolt Plus gels, LDS-PAGE and blotted for 70 minutes to PVDF using tank transfer. Blots were blocked immediately following transfer in 2% blocking reagent (GE RPN 2125; Healthcare) or 5% non-fat milk dissolved in 20 mM Tris, 137 mM sodium chloride pH 7.6 with 0.1% (v/v) Tween-20 (TBS-T) for 1h at room temperature with agitation. Blots were incubated in the primary antibody at a dilution of 1: 10 000 (in blocking reagent) for 1h at room temperature with agitation. The antibody solution was decanted and the blot was rinsed briefly twice, and then washed 1x15 min and 3x5 min with TBS-T at room temperature with agitation. Blots were incubated in secondary antibody (anti-rabbit IgG horse radish peroxidase conjugated, recommended secondary antibody AS09 602, Agrisera) diluted to 1:25 000 in blocking reagent for 1h at room temperature with agitation. The blots were washed as above. The blot was developed for 5 min with detection reagent according the manufacturers instructions. Images of the blots were obtained using a CCD imager (VersaDoc MP 4000) and Quantity One software (Bio-Rad). Exposure time was 10 seconds.Application examples: 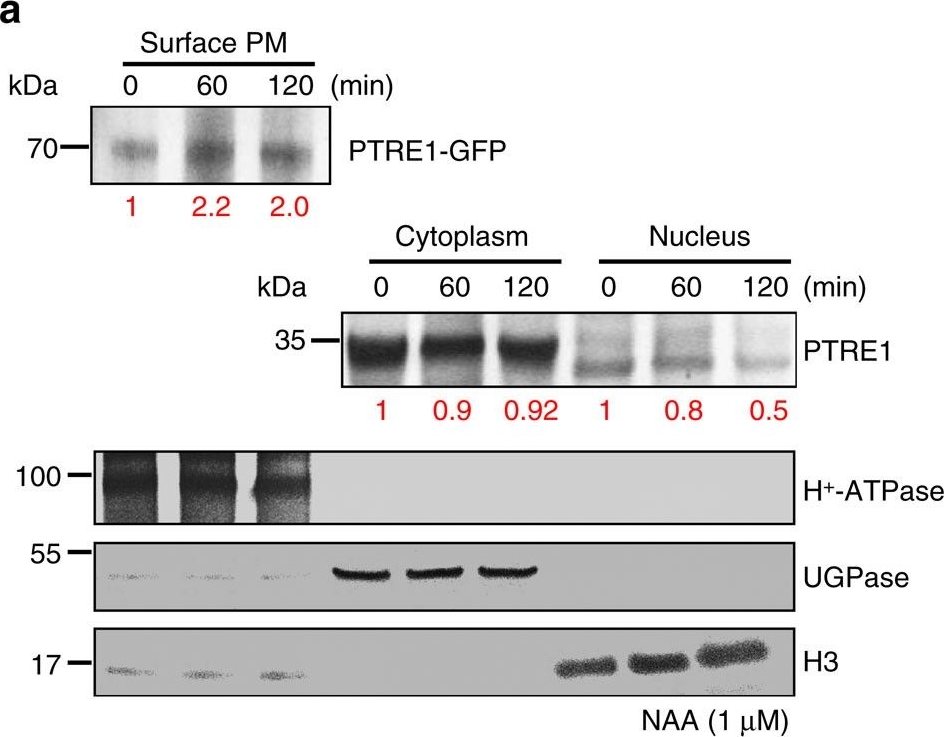
Reactant: Arabidopsis thaliana (Thale cress)
Application: Western Blotting
Pudmed ID: 27109828
Journal: Nat Commun
Figure Number: 6A
Published Date: 2016-04-25
First Author: Yang, B. J., Han, X. X., et al.
Impact Factor: 13.783
Open PublicationAuxin stimulates PTRE1 accumulation at the plasma membrane.(a) Western blot analysis showed that auxin treatment results in relatively more PTRE1 at the plasma membrane, but decreased levels in nucleus and cytoplasm. Arabidopsis seedlings expressing PTRE1-GFP were treated with NAA (1??M, 60 or 120?min) and then surface-exposed membrane proteins were analysed using PTRE1 antibody or plasma membrane marker H+-ATPase. Nucleus and cytoplasm fractions were prepared from Col seedlings and analysed by western blot using antibody against PTRE1, nuclear marker H3 or cytoplasm marker UGPase. The relative quantities of the proteins were calculated by using image pro plus and indicated. (b) A proposed model for how PTRE1 and TIR1 coordinate auxin responses regulating proteasome activity and Aux/IAA protein degradation. Under normal condition (with basal auxin levels), 26S proteasome activity is maintained by appropriate distribution of PTRE1 at the plasma membrane and in intracellular compartments. In response to auxin, auxin rapidly stimulates the association of TIR1 and Aux/IAA proteins resulting in degradation of Aux/IAAs (1). Later, auxin suppresses PTRE1 to inhibit proteasome activity (possibly through stimulating the accumulation of PTRE1 at plasma membrane, resulting in decreased intracellular and nuclear localization) and hence suppresses Aux/IAA protein degradation (2) to coordinate regulation of auxin responses, reflecting a mechanism for fine control of Aux/IAA homeostasis and auxin signalling.
- Additional Information
-
Additional information: Cellular [compartment marker] of cytoplasm, UGPse is a cytoplasmic protein Martz et al, (2002) Additional information (application): This antibody is also recognizing recombinant UGPase, below 0,5 pmol - Background
-
Background: UDP-glucose pyrophosphorylase (UGPase, UDPGP) E.C=2.7.7.9. is a key enzyme of synthesis of sucrose, cellulose and other saccharides. There are two cytoplasmic isoforms of UGPase-A (which share 94 % identity on amino acid level) and one chloroplastic UGPase-B isoform in Arabidopsis thaliana which share ca. 10-11 % of identity (Kleczkowski et al. 2011).
Alternative name: UTP--glucose-1-phosphate uridylyltransferase. - Product Citations
-
Selected references: Woo et al. (2026). TOE1 Interacts with FLM and SVP to Repress Flowering in Arabidopsis. J. of Plant Biology, Volume 69, pages 63–73.
Carrera-Castaño et al. (2026). Membrane-associated DELLA degradation modulates growth under carbon/nitrogen imbalance. Plant Cell. 2026 Jan 24:koag013. doi: 10.1093/plcell/koag013.
Martín-Merchán et al. (2024). Arabidopsis AGO1 N-terminal extension acts as an essential hub for PRMT5 interaction and post-translational modifications. Nucleic Acids Res . 2024 May 20:gkae387.doi: 10.1093/nar/gkae387.
Kleczkowski LA & Decker DD (2015) Sugar activation for production of nucleotide sugars as substrates for glycosyltransferases in plants. J. Appl. Glycosci. (in press). - Reviews:
-
This product doesn't have any reviews.