1
Anti-ACO1 | Aconitase
AS09 521 | Clonality:Polyclonal | Host:Rabbit | Reactivity: Arabidopsis thaliana, Solanum lycopersicum
Antibody replaced by AS23 4993
- Product Info
-
Immunogen: Arabidopsis ACO1 (AT4G35830, Q42560), codon 120 – 898 (C-terminus), was cloned in fusion with a N-terminal 6xHis tag, and over-expressed in E. coli. All recombinant protein accumulated in inclusion bodies, which were purified by centrifugation and solubilised in 6 M guanidine-HCl. The protein was refolded by dilution in 100 mM Tris-HCl 8.5, 10% (v/v) glycerol, 2 mM dithiothreitol, and concentrated prior to immunisation.
Host: Rabbit Clonality: Polyclonal Purity: Serum Format: Lyophilized Quantity: 100 µl Reconstitution: For reconstitution add 100 µl of sterile water Storage: Store lyophilized/reconstituted at -20°C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Western blot (WB) Recommended dilution: 1 : 5 000 -1 : 10 000 (WB), At higher concentrations the antibody binds aspecifically resulting in non-specific signals around 60 kDa, including Rubisco subunits Expected | apparent MW: 98 kDa, Note that ACO1, ACO2 and ACO3 cannot be distinguished in size by standard SDS-PAGE, - Reactivity
-
Confirmed reactivity: Arabidopsis thaliana ACO1,ACO2 and ACO3 isoforms, Brassica oleracea, Solanum lycopersicum Predicted reactivity: Cucurbita maxima, Hordeum vulgare, Nicotiana tabacum, Ricinus communis, Solanum tuberosum, Vitis vinifera, Oryza sativa, Zea mays, Picea sitcHensis, Populus trichocarpa
Species of your interest not listed? Contact usNot reactive in: Chlamydomonas reinhardii, cyanobacteria
- Application Examples
-
application example
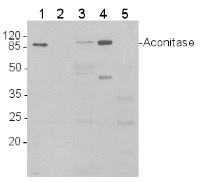
Western blot analysis of 1) 5 ng purified 6xHis-AtACO1 (Δ119, 87 kDa); 2) 2 ng 6xHis-AtACO1; 3) Total protein (15 μg) from Arabidopsis thaliana leaves were extracted with 2 volumes 50 mM Tris-HCl pH 8.0, 5% (v/v) glycerol, 1% (w/v) sodium dodecyl sulphate, 10 mM NaEDTA, 1 mM phenylmethanesulfonyl fluoride; 4) 15 µg of purified mitochondria from Arabidopsis thaliana cell culture, 5) 15 µg of protein from Arabidopsis thaliana chloroplasts
Proteins were separated by SDS-PAGE and blotted onto nitrocellulose (Whatman Protran BA 83, 0.2 μm). Blots were blocked in Tris-buffered saline (TBS) with 0.1% (v/v) Tween 20 and 5% (w/v) dried skimmed milk for 1 h at room temperature, and incubated with anti-AtACO1 antibodies diluted 1:10,000 in fresh block solution (10 mL per 8 x 6 cm blot) for 2 h at room temperature. The blot was washed 3 times with block solution, then incubated with horse-radish peroxidase conjugated anti-rabbit IgG antibodies, diluted 1: 5,000 in block solution, for 45 minutes. The blot was washed 2 times with block solution and 2 times with TBS-Tween. The signal was developed with standard ECL reagents and Kodak X-Omat LS film.
Note: as visible in lane 2, detection of recombinant AtACO1 falls below 2 ng of recombinant protein.
.jpg)
- Additional Information
-
Additional information: Arabidopsis expresses three highly similar aconitase isozymes (ACO1/ AT4G35830, ACO2/AT4G26970 and ACO3/AT2G05710), of which ACO1 is the cytosolic isoform, while ACO2 and ACO3 are predominantly located in the mitochondria (Arnaud et al 2007, Bernard et al 2009), The combined abundance and activity of the mitochondrial aconitases is about 3 times higher than the cytosolic pool (Bernard et al 2009),The Arabidopsis isoforms are more similar in amino acid sequence to mammalian iron-regulatory proteins (IRP-1) than to the mammalian and yeast mitochondrial aconitases Additional information (application): The antibody recognises all three Arabidopsis aconitase isoforms (ACO1, ACO2 and ACO3, see Bernard et al 2009), Possible differences in affinity have not been precisely quantified, Sensitivity threshold is between 2 and 10 ng for WB / ECL (see figure), Antibodies will recognize aconitase isoforms in denaturing and native gel electrophoresis - Background
-
Background: Aconitase is a single subunit enzyme of the tricarboxylic acid cycle (or Krebs cycle) in the mitochondria. A cytosolic isoform is also part of the glyoxylate cycle. Aconitase catalyzes the dehydration / hydration of citrate to iso-citrate, via cis-aconitate as an intermediate. The reaction is facilitated by an iron-sulphur cluster in the active site of the enzyme. The iron-sulphur cluster is somewhat unstable, especially under oxidative stress, and loss of the cofactor leads to degradation of the protein.Alternative names: ACO, citrate hydro-lyase 1,2,3
- Product Citations
-
Selected references: Pascual et al (2021). ACONITASE 3 is part of the ANAC017 transcription factor-dependent mitochondrial dysfunction response, Plant Physiology, 2021;, kiab225, https://doi.org/10.1093/plphys/kiab225
Przybyla-Toscano et al. (2021) Protein lipoylation in mitochondria requires Fe-S cluster assembly factors NFU4 and NFU5. Plant Physiol. 2021 Oct 28:kiab501. doi: 10.1093/plphys/kiab501. Epub ahead of print. PMID: 34718778.
Rurek et al. (2018). Mitochondrial Biogenesis in Diverse Cauliflower Cultivars under Mild and Severe Drought Involves Impaired Coordination of Transcriptomic and Proteomic Response and Regulation of Various Multifunctional Proteins. Preprints 2018, 2018010276 (doi: 10.20944/preprints201801.0276.v1).
Setién et al. (2014). Root phosphoenolpyruvate carboxylase and NAD-malic enzymes activity increase the ammonium-assimilating capacity in tomato. J Plant Physiol. 171:49-63.
Birke et al. (2012). Cysteine biosynthesis, in concert with a novel mechanism, contributes to sulfide detoxification in mitochondria of Arabidopsis thaliana. Biochem J. May 2, ahead of print.
- Protocols
-
Agrisera Western Blot protocol and video tutorials
Protocols to work with plant and algal protein extracts
Oxygenic photosynthesis poster by prof. Govindjee and Dr. Shevela
Z-scheme of photosynthetic electron transport by prof. Govindjee and Dr. Björn and Dr. Shevela - Reviews:
-
Michal RUREK | 2014-02-1410 µg of mitochondrial proteins from cauliflower (Brassica oleracea var. botrytis) curds (apical layer) was separated on 12 % SDS-PAGE (Laemmli- type) and blotted 1h to Immobilone P (Millipore) using Sedryt apparatus (Kucharczyk). After blocking (5% milk in PBS-T) for 1 h at RT, blots were incubated in the primary antibody at a dilution of 1: 10000 for 1h at room temperature with agitation. The primary antibody, diluted in 2% milk in PBS-T was reused several times. Secondary, anti-rabbit HRP- linked antibodies were bound at 1/10000 dilution. Single band of ca. 100 kDa was detected using standart GE Healthcare ECL reagents.


