1

Anti-ADGP | ADP-glucose pyrophosphorylase
AS11 1739 | Clonality: Polyclonal | Host: Rabbit | Reactivity: A. thaliana, C. reinhardtii, H. vulgare, N. tabacum, Polytomella sp., T. aestivum, Z. mays
- Product Info
-
Immunogen: Part of Arabidopsis thaliana recombinant ADP-glucose pyrophosphorylase small subunit, TAIR: At5g48300, UniProt: P55228
Host: Rabbit Clonality: Polyclonal Purity: Serum Format: Lyophilized Quantity: 50 µl Reconstitution: For reconstitution add 50 µl of sterile water Storage: Store lyophilized/reconstituted at -20°C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Western blot (WB) Recommended dilution: 1 : 1000-1 : 5000 (WB) Expected | apparent MW: 49.4 | 52 kDa - Reactivity
-
Confirmed reactivity: Arabidopsis thaliana, Horderum vulgare, Nicotiana tabacum, Triticum aestivum, Zea mays
Predicted reactivity: Chlamydiae, Chlamydomonas reinhardtii, Cyanobacteria, Beta vulgaris, Oryza sativa, Phaseolus vulgaris, Pisum sativum, Planctomycetes, Proteobacteria, Solanum lycopersicum, Spirochetes, Verrucomicrobia, Vitis vinifera Not reactive in: No confirmed exceptions from predicted reactivity are currently known - Application Examples
-
application example
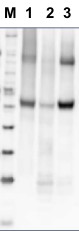
10 µg of total protein from Arabidopsis thaliana (1), Hordeum vulgare (2), Zea mays (3) total protein from all the samples were extracted with Protein Extraction Buffer PEB (AS08 300). Samples were diluted with 1X sample buffer (NuPAGE LDS sample buffer (Invitrogen) supplemented with 50 mM DTT and heat at 70 C for 5 min and keep on ice before loading. Protein samples were separated on NuPAGE 4-12% Tris-Bis gel (Invitrogen) LDS-PAGE and blotted for 1h to 1.5h on PVDF using tank transfer. Blots were blocked immediately following transfer in 2% blocking reagent (GE RPN 2125; Healthcare) or 5% non-fat milk dissolved in 20 mM Tris, 137 mM sodium chloride pH 7.6 with 0.1% (v/v) Tween-20 (TBS-T) for 1h at room temperature with agitation. Blots were incubated in the primary antibody at a dilution of 1: 5 000 (in blocking reagent) for 1h at room temperature with agitation. The antibody solution was decanted and the blot was rinsed briefly twice, and then washed 1x15 min and 3x5 min with TBS-T at room temperature with agitation. Blots were incubated in secondary antibody (anti-rabbit IgG horse radish peroxidase conjugated, recommended secondary antibody AS09 602) diluted to 1:25 000 in blocking reagent for 1h at room temperature with agitation. The blots were washed as above. The blot was developed for 5 min with TMA-6 (Lumigen) detection reagent according the manufacturers instructions. Images of the blots were obtained using a CCD imager (FluorSMax, Bio-Rad) and Quantity One software (Bio-Rad).
WT-Col-0 Arabidopsis thaliana leaves were frozen in liquid N2 and soluble proteins were extracted in a buffer containing 50 mM HEPES, 5 mM NaCl and 10 mM MgCl2. 10 µg and 17 ug of native protein complexes were separated by non-reducing blue native PAGE (5-12,5 %) and then further separated in a second dimension by reducing SDS-PAGE (15% with 6M urea) and blotted 1h to a PVDF membrane using Höefer semi-dry blotter. Blots were blocked with 2 % milk in TTBS for 3h at room temperature (RT) with agitation. Blot was incubated in the primary antibody at a dilution of 1: 2 000 overnight in +4°C. The antibody solution was decanted and the blot was washed 3 x 5 min with TTBS at RT with agitation. Blot was incubated in secondary antibody (anti-rabbit IgG horse radish peroxidase conjugated, from Agrisera AS09 602) diluted to 1:25 000 in 1% milk/TTBS for 2h at RT with agitation. The blot was washed 3x5 minutes in TTBS and 1x5 min in TBS at RT with agigation and developed for 5 min with ECL according to the manufacturer's instructions. Exposure time was 30 seconds.
2D gel electrophoresis was performed to exclude cross-reactivity of this antibody to Rubisco
Courtesy of Lauri Nikkanen, University of Turku, Finland
western blot on algal samples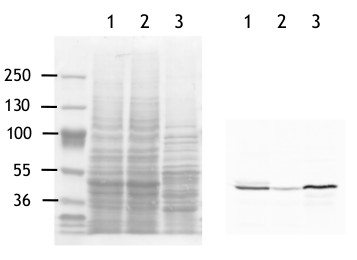
Total proteins (40 µg) from Chlamydomonas reinhardtii strain CC-124 (1), Chlamydomonas reinhardtii strain CC43-48 (STA6) (2) and Polytomella sp. Pringsheim 198.80 (3) were separated on a 5-12% acrylamide/6M urea SDS-PAGE and blotted onto nitrocellulose membrane. Blots were blocked in T-TBS (0.1% Tween in Tris Buffer Saline) containing 5% low fat milk, 1 h at room temperature (RT) with agitation. Blot was incubated in the primary antibody at a dilution of 1: 2 500 for 1h at RT with agitation. The antibody solution was decanted and the blot was washed twice for 15 min in T-TBS, RT with agitation. Blot was incubated in secondary antibody diluted to 1: 25 000 (goat anti-rabbit IgG, horse radish peroxidase conjugated, Agrisera, AS09 602), 1h at RT with agitation. The blot was washed as above and developed using the homemade enhanced chemiluminescence system (Durrant, Nature 346: 297, 1990). Images of the blots were obtained using a CCD imager (ImageQuant LAS4000).
Courtesy of Ariane Atteia, CNRS/Aix-Marseille Université, FranceApplication examples: 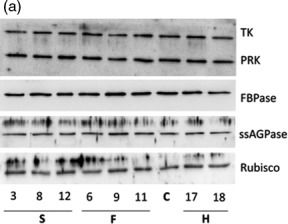
Reactant: Arabidopsis thaliana (Thale cress)
Application: Western Blotting
Pudmed ID: 27936496
Journal: Plant Biotechnol J
Figure Number: 2A
Published Date: 2017-07-01
First Author: Simkin, A. J., López-Calcagno, P. E., et al.
Impact Factor: 8.66
Open PublicationMolecular and biochemical analysis of the transgenic plants over?expressing SBPase (S), FBPA (F) and GDC?H (H). Immunoblot blot analysis of protein extracts from two independent leaves of (a) S, F and H lines and (b) SF and SFH lines used in this study compared to C. Transketolase (TK), phosphoribulokinase (PRK), fructose?1,6?bisphosphase (FBPase) the small subunit of ADP glucose pyrophosphoryalse (ssAGPase) and Rubisco.
Reactant: Arabidopsis thaliana (Thale cress)
Application: Western Blotting
Pudmed ID: 27936496
Journal: Plant Biotechnol J
Figure Number: 2B
Published Date: 2017-07-01
First Author: Simkin, A. J., López-Calcagno, P. E., et al.
Impact Factor: 8.66
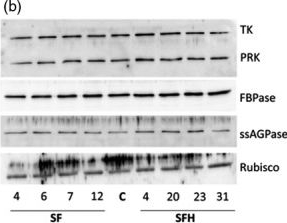
Open PublicationMolecular and biochemical analysis of the transgenic plants over?expressing SBPase (S), FBPA (F) and GDC?H (H). Immunoblot blot analysis of protein extracts from two independent leaves of (a) S, F and H lines and (b) SF and SFH lines used in this study compared to C. Transketolase (TK), phosphoribulokinase (PRK), fructose?1,6?bisphosphase (FBPase) the small subunit of ADP glucose pyrophosphoryalse (ssAGPase) and Rubisco.
- Additional Information
-
Additional information (application): Cross reactivity of this antibody to Rubisco has been excluded using 2D gel electrophoresis.
Antigen used to elicit this antbody has very poor conservation in large subunit of ADP-glucose pyrophosphorylase. - Background
-
Background: ADP-glucose pyrophosphorylase (AGPase) catalyses the first committed step in the synthesis of transient starch in chloroplasts and storage starch in amyloplasts in plants (Tetlow et al., 2004). AGPase in higher plants is composed of two distinct subunits encoded by separate genes, forming an L2S2 heterotetramer. The large subunits (L) are modulators of allosteric activity, while the small subunits (S) are the catalytic subunits (Kim et al., 2007). Synonymes:ADP-glucose pyrophosphorylase, ADP-glucose synthase, AGPase B, Alpha-D-glucose-1-phosphate adenyl transferase
- Product Citations
-
Selected references: Lim et al (2022). Arabidopsis guard cell chloroplasts import cytosolic ATP for starch turnover and stomatal opening. Nat Commun. 2022 Feb 3;13(1):652. doi: 10.1038/s41467-022-28263-2. PMID: 35115512; PMCID: PMC8814037.
Ma et al. (2021) A plasma membrane transporter coordinates phosphate reallocation and grain filling in cereals. Nat Genet. 2021 Apr 29. doi: 10.1038/s41588-021-00855-6. Epub ahead of print. PMID: 33927398.
Chang et al. (2020). Enhanced lipid productivity in AGP knockout marine microalga Tetraselmis sp. using a DNA-free CRISPR-Cas9 RNP method. Bioresource Technology.Volume 303, May 2020, 122932.
Ancin et al. (2019). NTRC and Thioredoxin f Overexpression Differentially Induces Starch Accumulation in Tobacco Leaves.Plants (Basel). 2019 Nov 26;8(12). pii: E543. doi: 10.3390/plants8120543.
Takahashi et al. (2019). Glutelin subtype-dependent protein localization in rice grain evidenced by immunodetection analyses. Plant Mol Biol. 2019 Mar 25. doi: 10.1007/s11103-019-00855-5.
Deng et al. (2018). Comparative Proteome Analysis of Wheat Flag Leaves and Developing Grains Under Water Deficit. Front Plant Sci. 2018 Apr 10;9:425. doi: 10.3389/fpls.2018.00425. eCollection 2018.
Yoshida et al. (2018). Thioredoxin-like2/2-Cys peroxiredoxin redox cascade supports oxidative thiol modulation in chloroplasts. Proc Natl Acad Sci U S A. 2018 Aug 28;115(35):E8296-E8304. doi: 10.1073/pnas.1808284115.
Zhen et al. (2018). 2D-DIGE comparative proteomic analysis of developing wheat grains under high-nitrogen fertilization revealed key differentially accumulated proteins that promote storage protein and starch biosyntheses. Anal Bioanal Chem. 2018 Jul 30. doi: 10.1007/s00216-018-1230-4. - Reviews:
-
Boon Leong Lim | 2022-04-14We used this antibodies (1:3000 dilution) to compare AGPase abundance in Arabidopsis mesophyll protoplasts and guard cell protoplasts. The results are satisfactory. Please refer tohttps://www.nature.com/articles/s41467-022-28263-2Miriam Schulz-Raffelt | 2021-06-23We used the ADGPase antibody in western blot application (dilution 1:2000) with total protein samples from Chlamydomonas reinhardtii. So far we got clear bands for a screening purpose, which was very satisfactory. Unfortunately, it seems that the frozen antibody in 1x PBS-TM cannot be reused because of high background.


