1
Anti-E(z) | Histone-lysine N-methyltransferase E(z)
AS16 3935 | Clonality: Polyclonal | Host: Rabbit | Reactivity: Drosophila melanogaster
- Product Info
-
Immunogen: GST-fusion with amino acids 8-190 of the Drosophila melanogaster E(z) protein, UniProt: P42124 Host: Rabbit Clonality: Polyclonal Purity: Immunogen affinity purified and depleted of anti-GST antibodies, in PBS pH 7.4. Format: Lyophilized Quantity: 50 µg Reconstitution: For reconstitution add 50 µl of sterile water Storage: Lyophilized antibody can be stored at -20°C for up to 3 years. Re-constituted antibody can be stored at 4°C for several days to weeks. Once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Chromatin Immunoprecipitation (IP) (ChIP) Recommended dilution: 3 µg/IP (ChIP) Expected | apparent MW: 86.9 kDa - Reactivity
-
Confirmed reactivity: Drosophila melanogaster
Predicted reactivity: Bactrocera cucurbitae, Bactrocera dorsalis, Bactrocera latifrons, Ceratitis capitata, Lucilia cuprina - Application Examples
-
application example
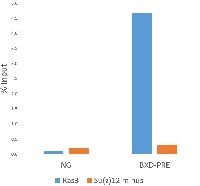
Figure 1. ChIP recovery.
ChIP and qPCR analysis were done as described [Schwartz YB, Kahn TG, Nix DA, Li XY, Bourgon R, et al. (2006) Genome-wide analysis of Polycomb targets in Drosophila melanogaster. Nat Genet 38: 700–705. doi: 10.1038/ng1817]. Chromatin from Su(z)12 mutant cell line (Kahn T. et all, submitted) served as a negative control, chromatin from Ras3 cells served as a positive control. Quantitative PCR was performed with primers specific for BXD-PRE of Ubx gene (Polycomb target gene in repressed state), used as positive controls, and for intergenic region, used as negative control. Figure 1 shows the ChIP recovery, measured by qPCR as a % of input (the relative amount of immunoprecipitated DNA compared to input DNA). Chromatin from 5x107 cells and 3mg of anti-E(z) antibody were used for each ChIP reaction.
Courtesy of Dr. Tatyana Khan, Umeå University, Sweden.
- Background
-
Background: Histone-lysine N-methyltransferase (Ez) is a Polycomb group (PcG) protein, and a catalytic subunit of the Esc/E(z) complex. By methylating Lys-9 and Lys-27 of histone H3, this complex is involved in transcriptional repression of target genes. E(z) is important for the repression during the first six hours of embryogenesis. The Esc/E(z) complex, together with the recruitment of the PRC1 complex, is necessary for the repression of homeotic target genes.
Alternative names: Lysine N-methyltransferase 6, Protein enhancer of zeste - Reviews:
-
This product doesn't have any reviews.

.JPG)

