1
PsbA | D1 protein of PSII positive control/quantitation standard
AS01 016S | Positive control/quantitation standard, for use in Western blot.
- Product Info
-
Format: Lyophilized in glycerol. Quantity: 100 µl Reconstitution: For reconstitution, check the label on the tube. Please notice that this product contains 10% glycerol and might appear as liquid but is provided lyophilized Storage: Store lyophilized/reconstituted at -20°C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Western blot (WB) Recommended dilution: Standard curve: 3 loads are recommended (0.5, 2 and 4 μl).
For most applications a sample load of 0.2 μg of chlorophyll will give a PsbA signal in this range.
Positive control: a 2 μl load per well is optimal for most chemiluminescent detection systems.
Non-disulphie dependent dimers and complexes can be also detected using standard western blot methods with more sensitive detection reagents as ECL Advance or West Pico when loading per well more standard than recommended. They have not been included in the standard calibration.
This standard is stabilized and ready and does not require heating before loading on the gel.
Please note that this product contains 10% glycerol and might appear as liquid but is provided lyophilized. Allow the product several minutes to solubilize after adding water. Mix thoroughly but gently Take extra care to mix thoroughly before each use, as the proteins tend to settle with the more dense layer after freezing.Expected | apparent MW: The standard has an actual MW of 41.5 kDa, The presence of a His6 tag causes it to run ~1.7 kDa higher on the gel than the native protein, Note that in most systems, PsbA migrates with an apparent MW of between 30 and 35 kDa. - Application Examples
-
Application example
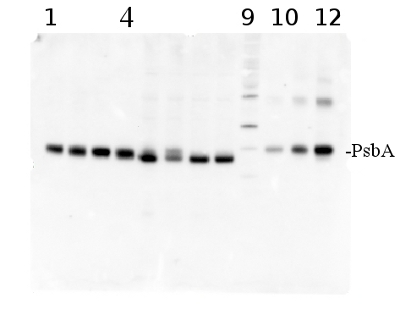
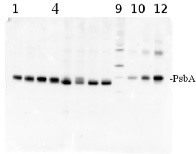
total protein from Synechococcus elongatus PCC 7942 (1-4) and Anabaena sp. PCC 7120 (5-8). Molecular weight markers (MagicMark XP, Invitrogen) (9). Recombinant PsbA protein standard (AS01 016S) is loaded in lanes 10-12 at 0.05 pmoles, 0.15 pmoles and 0.45 pmoles. Samples were separated on 4-12% NuPage (Invitrogen) LDS-PAGE and blotted 1h to PVDF. Blots were blocked immediately following transfer in 2% blocking reagent in 20 mM Tris, 137 mM sodium chloride pH 7.6 with 0.1% (v/v) Tween-20 (TBS-T) for 1h at room temperature with agitation. Blots were incubated in the primary antibody at a dilution of 1: 50 000 for 1h at room temperature with agitation. The antibody solution was decanted and the blot was rinsed briefly twice, then washed once for 15 min and 3 times for 5 min in TBS-T at room temperature with agitation. Blots were incubated in secondary antibody (anti-hen IgY horse radish peroxidase conjugated) diluted to 1:50 000 in 2% blocking solution for 1h at room temperature with agitation. The blots were washed as above and developed for 5 min with chemiluminescence detection reagent according the manufacturers instructions. Images of the blots were obtained using a CCD imager (FluorSMax, Bio-Rad) and Quantity One software (Bio-Rad).
Note: Optimal quantitation is achieved using moderate sample loads per gel lane, generally 0.5 to 2.5 ug total protein, depending on the abundance of the target protein.Quantitation: When quantitated standards are included on the blot, the samples can be quantitated using the available software. Excellent quantitation can be obtained with images captured on the Bio-Rad Fluor-S-Max or equivalent instrument using Bio-Rad QuantityOne software. The contour tool is used to select the area for quantitation and the values are background subtracted to give an adjusted volume in counts for each standard and sample. Using above protocol linear standard curves are generated over 1-1.5 orders of magnitude range in target load. It is important to note that immunodetections usually show a strongly sigmoidal signal to load response curve, with a region of trace detection of low loads, a pseudolinear range and a region of saturated response with high loads. For immunoquantitation it is critical that the target proteins in the samples and the standard curve fall within the pseudolinear range. Our total detection range using this protocol spans over 2 orders of magnitude, but the quantifiable range is narrower.
Quantitative western blot: detailed method description.
- Additional Information
-
Additional information: The PsbA protein standard can be used in combination with global anti-PsbA antibodies to quantitate PsbA from a wide range of species. Global antibodies are raised against highly conserved amino acid sequences in the PsbA protein.
Quantitative western blot: detailed method description, video tutorialThe goals when doing quantitative work:
- The sample PsbA must fall somewhere between the upper and lower standard loads.
- There should be at least 3 points on the standard curve.
- if possible, try to make the entire range of the curve around one order of magnitude or less (as in the application example).
- if possible, load <5µg total sample protein.
1pmol of PsbA standard is a strong load for chemiluminescence, but may be appropriate for the less sensitive reagents, for example alkaline phosphatase.
Additional information (application): Concentration: after adding 95 µl of sterile milliQ water final concentration of the standard is 0.25 pmoles/µl
Protein standard buffer composition: Glycerol 10%, Tris Base 141 mM, Tris HCl 106 mM, LDS 2%, EDTA 0.51 mM, SERVA® Blue G250 0.22 mM, Phenol Red 0.175 mM, pH 8.5, 0.1mg/ml PefaBloc protease inhibitor (Roche), 50mM DTT.
This standard is ready-to-load and does not require any additions or heating. It needs to be fully thawed and thoroughly mixed prior to using. Avoid vigorous vortexing, as buffers contain detergent. Following mixing, briefly pulse in a microcentrifuge to collect material from cap.
This standard is stabilized and ready and does not require heating before loading on the gel.
Please note that this product contains 10% glycerol and might appear as liquid but is provided lyophilized. Allow the product several minutes to solubilize after adding water. Mix thoroughly but gently Take extra care to mix thoroughly before each use, as the proteins tend to settle with the more dense layer after freezing. - Background
-
Background: The psbA gene has been cloned from many species of plants, green algae, and cyanobacteria. The psbA gene is located in the chloroplast genome and encodes for the D1 protein, a core component of Photosystem II. PsbA/D1 is rapidly cycled under illumination in all oxygenic photobionts. Tracking PsbA pools using the Global PsbA antibody can show the functional content of Photosystem II in a wide range of samples.
This is a recombinant protein standard, source: Synechocystis PCC 6803. - Product Citations
-
Selected references: Pipitone et al. (2021). A multifaceted analysis reveals two distinct phases of chloroplast biogenesis during de-etiolation in Arabidopsis. Elife. 2021 Feb 25;10:e62709. doi: 10.7554/eLife.62709. PMID: 33629953; PMCID: PMC7906606.
Fernández-González et al. (2020). Effects of Temperature and Nutrient Supply on Resource Allocation, Photosynthetic Strategy, and Metabolic Rates of Synechococcus Sp . J Phycol . 2020 Mar 4. doi: 10.1111/jpy.12983.
Levitan et al. (2019). Structural and functional analyses of photosystem II in the marine diatom Phaeodactylum tricornutum. Proc Natl Acad Sci U S A. 2019 Aug 27;116(35):17316-17322. doi: 10.1073/pnas.1906726116.
Ryan-Keogh et al. (2018). Seasonal regulation of the coupling between photosynthetic electron transport and carbon fixation in the Southern Ocean. Limnology and Oceanography.
Yuan et al. (2018). Combined effects of ocean acidification and warming on physiological response of the diatom Thalassiosira pseudonana to light challenges. Mar Environ Res. 2018 Apr;135:63-69. doi: 10.1016/j.marenvres.2018.01.016.
Li et al. (2016). A Hard Day's Night: Diatoms Continue Recycling Photosystem II in the Dark. Front. Mar. Sci., 08 November 2016.
Vandenhecke et al. (2015). Changes in the Rubisco to photosystem ratio dominates photoacclimation across phytoplankton taxa. Photosynth Res. 2015 Apr 11.
Wu et al. (2014). Large centric diatoms allocate more cellular nitrogen to photosynthesis to counter slower RUBISCO turnover rates. Front. Mar. Sci., 09 December 2014 | doi: 10.3389/fmars.2014.00068.
Li et al. (2014). The nitrogen costs of photosynthesis in a diatom under current and future pCO2. New Phytol. 2014 Sep 25. doi: 10.1111/nph.13037. - Protocols
-
- Reviews:
-
Orly Levitan | 2009-05-13I like Agrisera product, they work well in my assay and I am planning to keep on using your products as long as my assay will require it. Not that this standard is not as stable as the NifH.
Related products
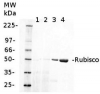
AS03 037 | Clonality: Polyclonal | Host: Rabbit | Reactivity: global antibody and compartment marker for higher plants, lichens, algae, cyanobacteria, dinoflagellates, diatoms
Benefits of using this antibody


