1
Anti-HSP70 | Heat shock protein 70 (cytoplasmic)
AS08 371 | Clonality: Polyclonal | Host: Rabbit | Reactivity: A. thaliana, B. juncea, B. oleracea, C. sativus, C. reinhardtii, D. subspicatus, E. tef, G. vermiculophylla, H. vulgare, M. sativa, O. sativa, P. kurroa, P. strobus, Salicornia sp., S. italica. S. vulgaris, S. lycopersicum, Trebouxia TR1 and TR9, T. aestivum, Z. mays, P. falciparum, V. faba
Benefits of using this antibody
- Product Info
-
Immunogen: KLH-conjugated synthetic peptide conserved in known higher plant HSC70 proteins including three isoforms of Arabidopsis thaliana HSC70-1 UniProt: F4KCE5 , HSC70-2 UniProt: A0A178UTH3 and HSC70-3 Uniprot: O65719 as well as heat shock inducible Hsp70 of Arabidopsis thaliana TAIR: AT3g12580/T2E22_110 and At1g16030 and AT3g12580/T2E22_110Host: Rabbit Clonality: Polyclonal Purity: Serum Format: Lyophilized Quantity: 50 µl Reconstitution: For reconstitution add 50 µl of sterile water Storage: Store lyophilized/reconstituted at -20°C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Immunoprecipitation (IP), Western blot (WB) Recommended dilution: 1 : 3000 - 1: 10 000, 5 µg protein/well (WB), 2-3 µl/protein extract of concentration 3-5 mg/ml Expected | apparent MW: 70 kDa
- Reactivity
-
Confirmed reactivity: Arabidopsis thaliana, Brassica juncea, Brassica napus, Brassica oleracea, Chlamydomonas reinhardtii, Chlamydomonas sp. UWO241, Cucumis sativus, E. teft, Fagopyrum esculentum, Glycine max, Hordeum vulgare, Medicago sativa, Olea europea, Picrorhiza kurroa, Pinus strobus, Polyscias elegans, Salicornia sp., Silene vulgaris, Solanum lycopersicum, Zea mays, algae: Desmodesmus subspicatus, Gracilaria vermiculophylla, phycobiont: Trebouxia TR1 and TR9, Plasmodium falciparum, Setaria italica, Triticum aestivum, Ulva prolifera, Vicia faba
Predicted reactivity: Ageratina adenophora, Allium sativum, Arabis alpina, Arachis diogoi, Arundo donax, Brassica napus, brassica rapa subsp. pekinensis, Camellia sinensis, Chlorella vulgaris, Citrus clementina, Citrus sinensis, Coffea arabica, Eriobotrya japonica, Gossypium arboretum, Glycine soja, Helianthus annuus, Hordeum vulgare var. distichum, Lolium multiflorum , Lotus japonicus, Medicago sativa, medicago truncatula, Musa acuminata subsp. malaccensis, Nannochloropsis gaditana, Nicotiana tabacum, Nicotiana bethamiana, Oryza sativa, Phaseolus vulgaris, Physcomitrium patens, Pinus taeda, Pisum sativum, Populus balsamifera, Populus trichocarpa, Saccharum sp, Salix gilgiana, Saussurea medusa,Solanum tuberosum, Solanum commersonii, Spinacia oleracea, Tragopogon dubius, Tragopogon porrifolius, Triticum aestivum, Vitis vinifera, Volvox sp.
Species of your interest not listed? Contact usNot reactive in: Centella asiatica, Polyscias elegans - Application Examples
-
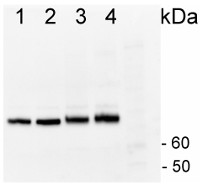
1µg of total protein from Horderum vulgare pre heat shock leaf (1), Horderum vulgare post heat shock (2h 40ºC) (2), Zea mays pre heat shock total protein leaf (3), Zea mays post heat shock (2h 40ºC) (4), total protein leaf extracted with Agrisera Protein Eextraction Buffer (AS08 300) were separated on 4-12% NuPage (Invitrogen) LDS-PAGE and blotted 1h to PVDF (Milipore). Filters were blocked 1h with 2% low-fat milk powder in TBS-T (0.1% TWEEN 20) and probed with anti-HSP70 antibody (AS08 371, 1:20 000, 1h) and secondary anti-rabbit (1:20 000, 1 h) antibody (HRP conjugated) in TBS-T containing 2% low fat milk powder. All steps were performed at RT with agitation. Signal was detected with chemiluminescent detection reagent with extreme femtogram range.
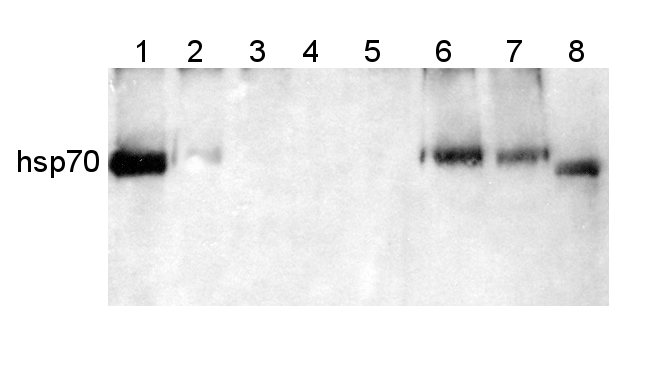
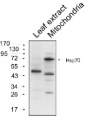
Protein from Solanum lycopersicum (1) total cell extract ca. 30 -50 µg, (2) and (3) nuclei pellet , (4) and (5) ca. 7 µg of nuclei fraction, (6) and (7) cytoplasmic pellet, (8) ca. 7 µg of cytoplasm fraction, were separated on 10% SDS-PAGE and blotted 1h to nitrocellulose (Schleicher & Schuell). Filters were blocked 1h with 2% low-fat milk powder in TBS-T (0.1% TWEEN 20) and probed with anti-HSP70 antibody (AS08 371, 1:5000, 3h RT). The antibody solution was decanted and the blot was rinsed briefly. Washed 3 times for 15 min in TBS-T at room temperature with agitation. Blot was incubated with a secondary antibody (anti-rabbit IgG horse radish peroxidase conjugated) diluted to 1: 5:000. The blot was washed as above and developed for 1 min with ECL detection reagent according to the manufacturers instructions.
Courtesy Dr Rena Gorovits, The Hebrew University of Jerusalem, Israel
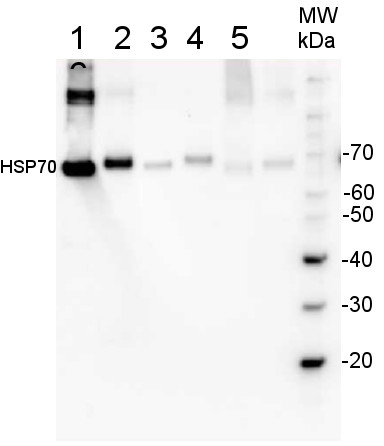
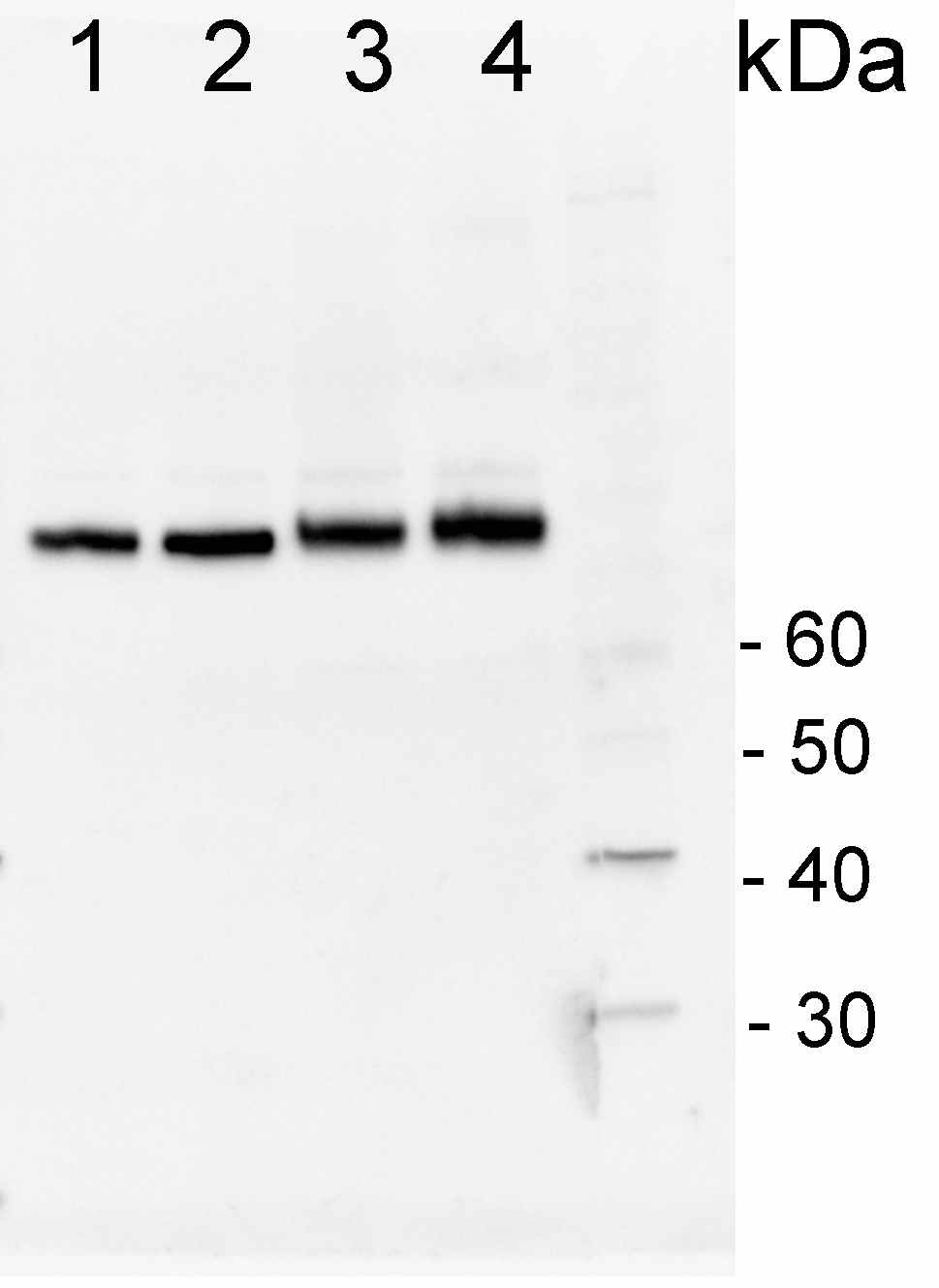
200 fmoles of HSP70 protein standard product number AS08 371S (1), 1 µg of total protein from samples such as Lycopersicum esculentum leaf (2), Nicotiana tabaccum leaf, (3), Zea mays leaf (4), Hordeum vulgare leaf (5), Arabidopsis thaliana leaf (6) were extracted with Agrisera Protein Extraction Buffer PEB (AS08 300). Samples were diluted with 1X sample buffer (NuPAGE LDS sample buffer (Invitrogen) supplemented with 50 mM DTT and heat at 70°C for 5 min and keept on ice before loading. Protein samples were separated on 4- 12% Bolt Plus gels, LDS-PAGE and blotted for 70 minutes to PVDF using tank transfer. Blots were blocked immediately following transfer in 2% blocking reagent or 5% non-fat milk dissolved in 20 mM Tris, 137 mM sodium chloride pH 7.6 with 0.1% (v/v) Tween-20 (TBS-T) for 1h at room temperature with agitation. Blots were incubated in the primary antibody at a dilution of 1: 10 000 (in blocking reagent) for 1h/RT with agitation. The antibody solution was decanted and the blot was rinsed briefly twice, and then washed 1x15 min and 3x5 min with TBS-T at room temperature with agitation. Blots were incubated in secondary antibody (anti-rabbit IgG horse radish peroxidase conjugated, recommended secondary antibody AS10 1489, Agrisera) diluted to 1:25 000 in blocking reagent for 1h at room temperature with agitation. The blots were washed as above. The blot was developed for 5 min with chemiluminescence detection reagent in extreme femtogram range, according the manufacturers instructions. Images of the blots were obtained using a CCD imager (VersaDoc MP 4000) and Quantity One software (Bio-Rad). Exposure time was 30 seconds.
Application examples: 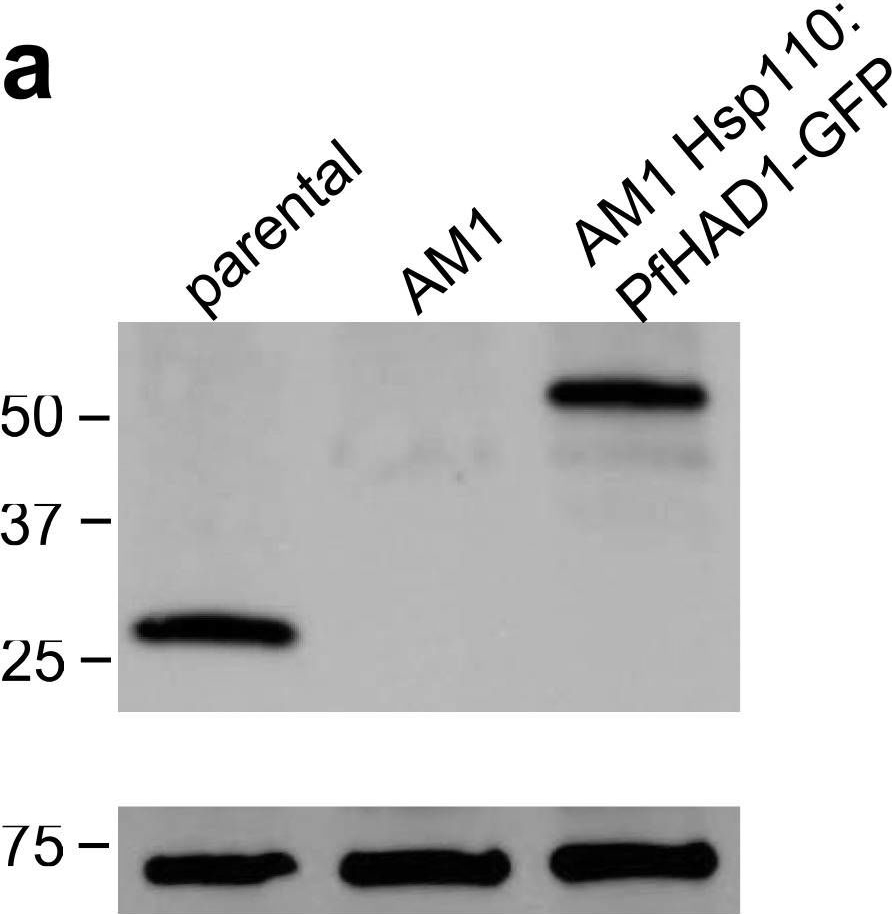
Reactant: Plasmodium falciparum
Application: Western Blotting
Pudmed ID: 25058848
Journal: Nat Commun
Figure Number: 5A
Published Date: 2014-07-24
First Author: Guggisberg, A. M., Park, J., et al.
Impact Factor: 13.783
Open PublicationLoss of PfHAD1 is required for FSM resistanceData shown are representative of at least three independent experiments. (a) Immunoblot of the parent strain, FSMR strain AM1, and AM1 Hsp110:PfHAD1-GFP. Full blots are shown in Supplementary Fig. 3. Marker units are kilodaltons (kDa). Top panel was probed with anti-PfHAD1 antisera, and the bottom panel was probed with anti-heat shock protein 70 (hsp70) antisera as a loading control. Expected approximate protein masses: native PfHAD1, 33 kDa; PfHAD1-GFP, 60 kDa; hsp70, 74 kDa. (b) FSM IC50s of the parent strain, FSMR strain AM1, and AM1 Hsp110:PfHAD1-GFP. Displayed are the means ± standard error of the mean (SEM). The parent strain has a FSM IC50 of 1.1 ± 0.2 ?M, while AM1 has an IC50 of 5.5 ± 1.5 ?M. Expression of PfHAD1-GFP in AM1 results in an IC50 of 1.4 ± 0.2 ?M. Data shown are mean is normalized such that 100% growth is defined by that of untreated cells, and 0% growth is defined as the smallest value in each data set.
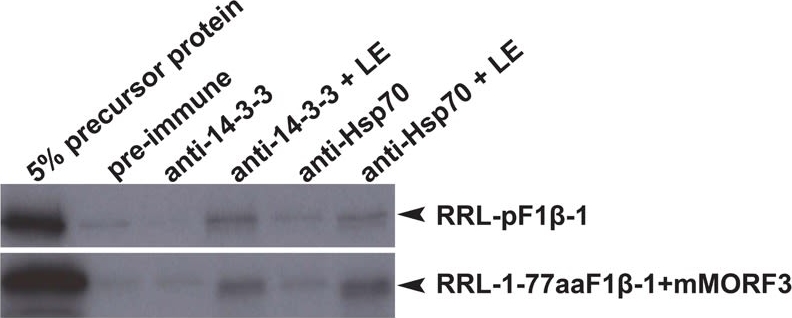
Reactant: Arabidopsis thaliana (Thale cress)
Application: Western Blotting
Pudmed ID: 30042778
Journal: Front Plant Sci
Figure Number: 4A
Published Date: 2018-07-26
First Author: Law, Y. S., Ngan, L., et al.
Impact Factor: 5.435
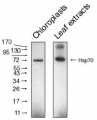
Open PublicationCo-immunoprecipitation assays of pF1?-1. The binding of RRL-pF1?-1 and RRL-1-77aaF1?-1 + mMORF3 to Hsp70 and 14-3-3 were strongly enhanced by LE treatment.
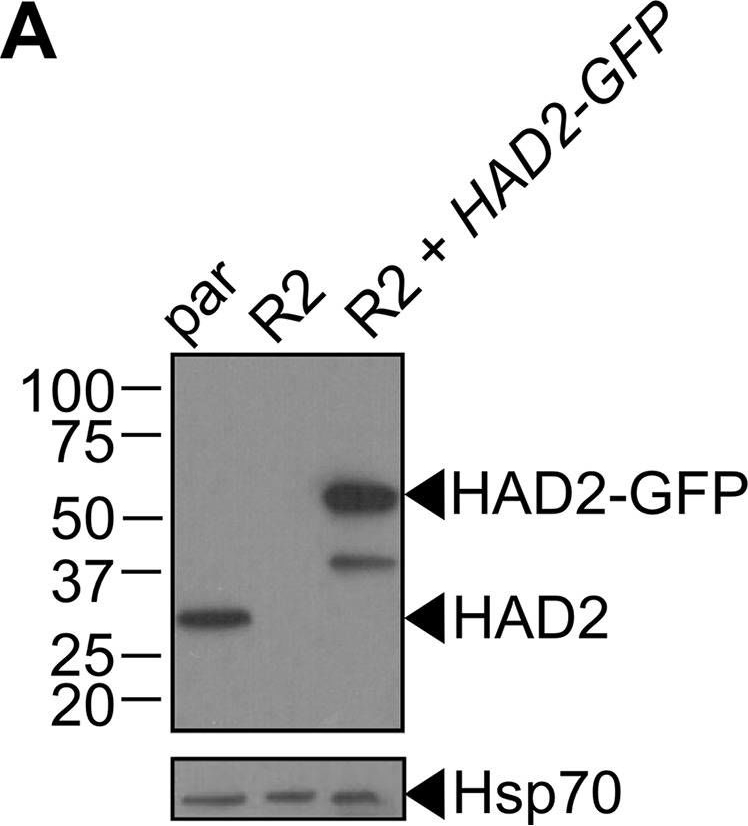
Reactant: Plasmodium falciparum
Application: Western Blotting
Pudmed ID: 30425143
Journal: MBio
Figure Number: 5A
Published Date: 2018-11-13
First Author: Guggisberg, A. M., Frasse, P. M., et al.
Impact Factor: 6.689
Open PublicationLoss of HAD2 is necessary for FSM resistance. (A) Successful expression of pTEOE110:HAD2-GFP in strain R2 (had2R157X, PFK9) was confirmed by immunoblotting. Marker units are indicated in kilodaltons (kDa). The top blot was probed with anti-HAD2 antiserum (expected masses: HAD, 33?kDa; HAD2-GFP, 60?kDa). The bottom blot was probed with anti-heat shock protein 70 (Hsp70) antiserum as a loading control. (B) Representative FSM dose response demonstrating that expression of HAD2-GFP in strain R2 (had2R157X PFK9) resulted in restored sensitivity to FSM. Strain R2 had an elevated FSM IC50 compared to the parental (par) strain. When HAD2 expression was restored in strain R2, the resulting strain showed an IC50 near that of the parent strain. Data shown are from a representative clone (clone 1) of the HAD2-rescued strain. Additional clones displayed a similar phenotype (see Fig. S2).
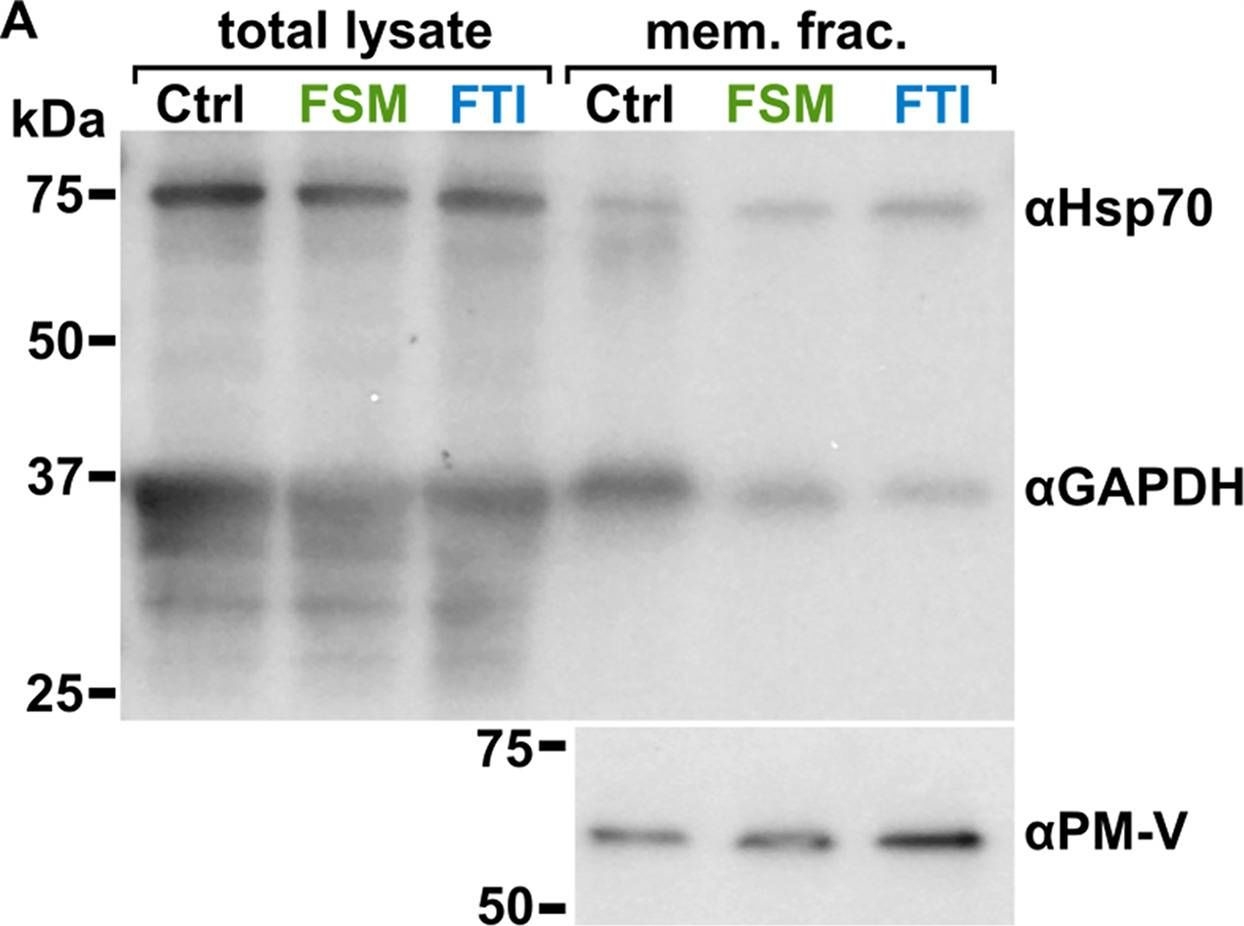
Reactant: Plasmodium falciparum
Application: Western Blotting
Pudmed ID: 34182772
Journal: MBio
Figure Number: 7A
Published Date: 2021-06-29
First Author: Mathews, E. S., Jezewski, A. J., et al.
Impact Factor: 6.689
Open PublicationLocalization of GAPDH, but not its glycolytic function, is IPP- or farnesylation-dependent. (A to C) Inhibition of IPP synthesis and farnesylation reduced membrane association of GAPDH. (A) Representative anti-GAPDH immunoblots of control-, FSM (20??M)-, and FTI (10??M)-treated P. falciparum. (B and C) Quantification of several immunoblots adjusted with loading control. Anti-Hsp70 (1:5,000) and anti-PM-V (1:500) were used as loading controls for total lysate and membrane fractions, respectively. n?=?5 to 6; ****, P???0.0001, unpaired t test with Welch’s correction.
- Additional Information
-
Additional information: This product can be sold containing ProClin if requested Additional information (application): Can be sold containing 0.1% ProClin if requested
This antibody can be used as a marker of cytoplasmic fraction in tomato (Anfoka et al. 2015).
Applied primary antibody dilution in western blot depends upon sensitivity of detection reagents (pico or femtogram for chemiluminescent detection).
Immunoprecipitation protocol using Agrisera anti-Hsp70 cytosolic antibodies, see tab: protocols. - Background
-
Background: Heat-shock protein 70 (Hsp70) is the major stress-inducible protein in vertebrates and is highly conserved throughout evolution. It plays a role as a molecular chaperone and is important for allowing cells to cope with acute stressor insult, especially those affecting the protein machinery. Heat shock cognate protein 70 (HSC70), is a highly conserved protein and a member of the family of molecular chaperones. - Product Citations
-
Selected references: Bělonožníková et al. (2026). Biochemical impacts of triazole fungicides on tomato roots and soil enzyme activities. J Environ Sci Health B . 2026 Mar 24:1-15. doi: 10.1080/03601234.2026.2644792.
Çetinbaş Genç et al. (2026). Tolerance mechanisms to UV-B in olive pollen demonstrate cultivar-dependent biochemical responses. Plant Physiol Biochem. 2026 Apr 2:233:111264. doi: 10.1016/j.plaphy.2026.111264.
Pühringer et al. (2025). Automated mass spectrometry-based profiling of multi-glycosylated glycosyl inositol phospho ceramides (GIPC) reveals specific series GIPC rearrangements during barley grain development and heat stress response. lant J . 2025 Jun;122(6):e70279. doi: 10.1111/tpj.70279.
Tanaka et al. (2024). Heat shock protein 70 is associated with duration of cell proliferation in early pod development of soybean. Commun Biol. 2024 Jun 21;7(1):755. doi: 10.1038/s42003-024-06443-8.
Vuković et al. (2024). Heat Priming Modifies Heat Stress Response in BPM1-Overexpressing Arabidopsis thaliana (L.) Heynh. J Olant Growth Regulation. https://doi.org/10.1007/s00344-024-11337-4.
Llamas et al. (2023). In planta expression of human polyQ-expanded huntingtin fragment reveals mechanisms to prevent disease-related protein aggregation. Nat Aging. 2023 Nov;3(11):1345-1357.doi: 10.1038/s43587-023-00502-1.
Tokić, M.; Leljak Levanić, D.; Ludwig-Müller, J.; Bauer, N. Growth and Molecular Responses of Tomato to Prolonged and Short-Term Heat Exposure. Int. J. Mol. Sci. 2023, 24, 4456.
Stachurska, J.; Sadura, I.; Rys, M.; Dziurka, M.; Janeczko, A. Insight into Hormonal Homeostasis and the Accumulation of Selected Heat Shock Proteins in Cold Acclimated and Deacclimated Winter Oilseed Rape (Brassica napus L.). Agriculture 2023, 13, 641.
Flores-Cáceres et al. (2023). The Early Oxidative Stress Induced by Mercury and Cadmium Is Modulated by Ethylene in Medicago sativa Seedlings. Antioxidants 2023, 12, 551.
Dong et al. (2023) ASYMMETRIC LEAVES 2 and ASYMMETRIC LEAVES 2-LIKE are partially redundant genes and essential for fruit development in tomato. Plant J. 2023 Jun;114(6):1285-1300. doi: 10.1111/tpj.16193
Cvetkovska et al. (2022) A constitutive stress response is a result of low temperature growth in the Antarctic green alga Chlamydomonas sp. UWO241. Plant, Cell & Environment, 45, 156– 177.
Belonoznikova et al. (2022). Seed Protection of Solanum lycopersicum with Pythium oligandrum against Alternaria brassicicola and Verticillium albo-atrum. Microorganisms. 2022 Jul 4;10(7):1348. doi: 10.3390/microorganisms10071348. PMID: 35889067; PMCID: PMC9315653.
Chong et al. (2022) The tomato OST1-VOZ1 module regulates drought-mediated flowering. Plant Cell. 2022 Apr 26;34(5):2001-2018. doi: 10.1093/plcell/koac026. PMID: 35099557; PMCID: PMC9048945.
Bychkov et al. (2022) The role of PAP4/FSD3 and PAP9/FSD2 in heat stress responses of chloroplast genes. Plant Sci. 2022 Sep;322:111359. doi: 10.1016/j.plantsci.2022.111359. Epub 2022 Jun 20. PMID: 35738478.
Cvetkovska et al. (2022) A constitutive stress response is a result of low temperature growth in the Antarctic green alga Chlamydomonas sp. UWO241. Plant, Cell & Environment, 45, 156– 177. https://doi.org/10.1111/pce.14203
Wang et al. (2022) 17-(Allylamino)-17-demethoxygeldanamycin treatment induces the accumulation of heat shock proteins and alleviates senescence in broccoli. Postharvest Biology and Technology,Volume 186, 2022, 111818, ISSN 0925-5214, https://doi.org/10.1016/j.postharvbio.2021.111818.
Kumari et al. (2021) In-depth assembly of organ and development dissected Picrorhiza kurroa proteome map using mass spectrometry. BMC Plant Biol. 2021 Dec 22;21(1):604. doi: 10.1186/s12870-021-03394-8. PMID: 34937558; PMCID: PMC8693493.
Mishra et al. (2021) Interplay between abiotic (drought) and biotic (virus) stresses in tomato plants. Mol Plant Pathol. 2021 Dec 30. doi: 10.1111/mpp.13172. Epub ahead of print. PMID: 34970822.
Shteinberg et al. (2021) Tomato Yellow Leaf Curl Virus (TYLCV) Promotes Plant Tolerance to Drought. Cells. 2021 Oct 25;10(11):2875. doi: 10.3390/cells10112875. PMID: 34831098; PMCID: PMC8616339.
Swetha et al. (2021) Single and Combined Salinity and Heat Stresses Impact Yield and Dead Pericarp Priming Activity. Plants (Basel). 2021 Aug 8;10(8):1627. doi: 10.3390/plants10081627. PMID: 34451672; PMCID: PMC8399105.
Plazek et al. (2020). Synthesis of heat-shock proteins HSP-70 and HSP-90 in flowers of common buckwheat (Fagopyrum esculentum) under thermal stress. Crop and Pasture Science, 71(8), 760-767, July 2020
Azaiez et al. (2020). Salt Stress Induces Differentiated Nitrogen Uptake and Antioxidant Responses in Two Contrasting Barley Landraces from MENA Region. Agronomy 10, no. 9: 1426.
Sadura et al. (2020). HSP Transcript and Protein Accumulation in Brassinosteroid Barley Mutants Acclimated to Low and High Temperatures . Int J Mol Sci . 2020 Mar 10;21(5):1889.doi: 10.3390/ijms21051889.
Tabassum et al. (2020). FLOURY ENDOSPERM11-2 Encodes Plastid HSP70-2 Involved With Temperature-Dependent Chalkiness of Rice (Oryza Sativa L.) Grains. Plant J. 10.1111/tpj.14752
Rowarth et al. (2019). Hsp70 plays a role in programmed cell death during the remodelling of leaves of the lace plant (Aponogeton madagascariensis). J Exp Bot. 2019 Nov 6. pii: erz447. doi: 10.1093/jxb/erz447
McLoughlin et al. (2019) HSP101 Interacts with the Proteasome and Promotes the Clearance of Ubiquitylated Protein Aggregates. Plant Physiol. 2019 Aug;180(4):1829-1847. doi: 10.1104/pp.19.00263
Deng et al. (2019). Integrated proteome analyses of wheat glume and awn reveal central drought response proteins under water deficit conditions. J Plant Physiol. 2019 Jan;232:270-283. doi: 10.1016/j.jplph.2018.11.011.
Lentini et al. (2018). Early responses to cadmium exposure in barley plants: effects on biometric and physiological parameters. Acta Physiologiae Plantarum October 2018, 40:178
Stachurska, J.; Sadura, I.; Rys, M.; Dziurka, M.; Janeczko, A. Insight into Hormonal Homeostasis and the Accumulation of Selected Heat Shock Proteins in Cold Acclimated and Deacclimated Winter Oilseed Rape (Brassica napus L.). Agriculture 2023, 13, 641.
Fan et al. (2018). Comparative proteomic analysis of Ulva prolifera response to high temperature stress. Proteome Sci. 2018 Oct 27;16:17. doi: 10.1186/s12953-018-0145-5.
Pan et al. (2018). Comparative proteomic investigation of drought responses in foxtail millet. BMC Plant Biol. 2018 Nov 29;18(1):315. doi: 10.1186/s12870-018-1533-9.
Lentini et al. (2018). Early responses to cadmium exposure in barley plants: effects on biometric and physiological parameters. Acta Physiol Plant (2018) 40: 178.
Balazova et al. (2018). Zinc oxide nanoparticles phytotoxicity on halophyte from genus Salicornia. Plant Physiol Biochem. 2018 Sep;130:30-42. doi: 10.1016/j.plaphy.2018.06.013.
Yoon et al. (2018). The subfamily II catalytic subunits of protein phosphatase 2A (PP2A) are involved in cortical microtubule organization. Planta. 2018 Sep 6. doi: 10.1007/s00425-018-3000-0.
Alamri et al. (2018). Nitric oxide-mediated cross-talk of proline and heat shock proteins induce thermotolerance in Vicia faba L. Environmental and Experimental Botany Available online 23 June 2018.
Barghetti et al. (2017). Heat-shock protein 40 is the key farnesylation target in meristem size control, abscisic acid signaling, and drought resistance. Genes Dev. 2017 Nov 15;31(22):2282-2295. doi: 10.1101/gad.301242.117.
Gorovits et al. (2017). The six Tomato yellow leaf curl virus genes expressed individually in tomato induce different levels of plant stress response attenuation. Cell Stress Chaperones. 2017 Mar 21. doi: 10.1007/s12192-017-0766-0.
Fernandez-Bautista N. et al. (2017). AtHOP3, a member of the HOP family in Arabidopsis, interacts with BiP and plays a major role in the ER stress response. Plant Cell Environ. 2017 Feb 2. doi: 10.1111/pce.12927.
Hammann et al. (2016). Selection of heat‑shock resistance traits during the invasion of the seaweed Gracilaria vermiculophylla. Marine Biology 163: 104.
McLoughlin et al. (2016) Class I and II Small Heat Shock Proteins Together with HSP101 Protect Protein Translation Factors during Heat Stress. Plant Physiol. 2016 Oct;172(2):1221-1236.
Shen et al. (2016). The Arabidopsis polyamine transporter LHR1/PUT3 modulates heat responsive gene expression by enhancing mRNA stability. Plant J. 2016 Aug 19. doi: 10.1111/tpj.13310. [Epub ahead of print]
Gorovits et al. (2016). Tomato yellow leaf curl virus confronts host degradation by sheltering in small/midsized protein aggregates. Virus Res. 2016 Feb 2;213:304-13. doi: 10.1016/j.virusres.2015.11.020. Epub 2015 Dec 1.
Ghandi et al. (2016). Tomato yellow leaf curl virus infection mitigates the heat stress response of plants grown at high temperature. Sci Rep. 2016 Jan 21;6:19715. doi: 10.1038/srep19715. - Protocols
-
Agrisera Western Blot protocol and video tutorials
Protocols to work with plant and algal protein extracts
Oxygenic photosynthesis poster by prof. Govindjee and Dr. Shevela
Z-scheme of photosynthetic electron transport by prof. Govindjee and Dr. Björn and Dr. ShevelaCo-Immunoprecipitaion using Agrisera HSP70 cytoplasmic antibody
2-3 µl of Agrisera anti-Hsp70 cytosolic antibody is added to ProteinA in IP buffer:
50 mM tris pH 7.5
100 mM KCl
10 mM MgCl2
rotation with antibody: 2-3 h at 4ºC
The unbound antibody must be washed in the same buffer 3-4 times
Protein extract with the concentration not lower then 3-5 mg/ml is added to ProteinA with anti-Hsp70 in amounts of 400-600 µl and is rotated for ON at 4ºC.
Unbound material is washed by IP buffer twice, then twice in the same IP buffer which contains 0.25 M NaCl.
All supernatant is carefully removed, SDS-PAGE sample buffer is added to the beads, and boiled for 10 min. After the centrifugation (12500 rpm 5 min) supernatant is loaded on a gel.Courtesy Dr. Rena Gorovits, The Hebrew University of Jerusalem, Israel
- Reviews:
-
Wendy | 2025-10-22Although the species we need to detect is not listed in the reactivity section on the website, we can still use it to test the new plant experiments.Alberto Dávalos | 2024-04-12This antibody worked very well. Dilution 1:2000. We propose as a marker of plant-derived extracellular vesicles. It works in A. thaliana, broccoli, pomegranate and orange. Pharmacol Res. 2023 Dec198:106999. doi: 10.1016/j.phrs.2023.106999.Shelby Hotz | 2023-03-21For my lab's study, we used this antibody for western blots to detect the abundance of Hsp70 in heat shocked and non-heat shocked bull kelp (Nereocystis leutkeana). We used a 1:3000 dilution in a 5% Nonfat Dried Milk in 1x Tris Buffered Saline solution. The results were positive, with the protein bands showing up clearly and at the correct weight according to the positive control we used.
I am happy to report the antibody has been working effectively and as intended in our molecular research. The instructions that come with the product are very comprehensive and relatively easy to follow, and doing so has allowed us to effectively use the materials as expected and needed. We also appreciate the promptness of the delivery and arrival of the materials, and having the antibody in a lyophilized form is very much appreciated.
Thank you for your service.?udmila Holubová | 2019-10-04We are very satisfied with this antibody. We used it in Zea mays protein extract in dilution 1:3000, but I agree, it could be used also in higher dilution.| 2011-10-04We tried with success this antibody in Medicago sativa, with 10 ug protein loading, with a 1/5000 dilution of primary antibody and 1/20000 secondary HRP, suitable for ECL detection. Due its high reactivity, a higher dilution of primary antibody could be tried.Dr. Rena Gorovits | 2009-04-30Excellent antibody distinguishes between cytoplasmic and nuclear fractions in tomato protein extracts.
Related products
AS08 347 | Clonality: Polyclonal | Host: Rabbit | Reactivity: Arabidopsis thalian, Brassica oleracea, Hordeum vulagre
AS08 348 | Clonality: Polyclonal | Host: Rabbit | Reactivity: A. thaliana, H. vulgare, O. sativa, P. sativm, P. strobus

