1
Anti-NR | Nitrate reductase, assimilatory
AS08 310 | Clonality: Polyclonal | Host: Rabbit | Reactivity: A. thaliana, H. vulgare, C. reinhardtii, red alga Gracilaria gracilis, Medicago sativa, diatom Thalassiosira sp., P. tricornutum Bohlin, P. yunnanensis Dode, P. notoginseng, S. lycopersicum, S. tuberosum
- Product Info
-
Immunogen: KLH-conjugated synthetic peptide derived from conserved domain in NADH-NR protein sequences including A.thaliana NR1 P11832, At1g77760 and NR2 P11035, At1g37130
Host: Rabbit Clonality: Polyclonal Purity: Immunogen affinity purified serum in PBS pH 7.4. Format: Lyophilized Quantity: 100 µg Reconstitution: For reconstitution add 50 µl of sterile water Storage: Store lyophilized/reconstituted at -20°C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Western blot (WB) Recommended dilution: 1 : 500 -1 : 1000 (WB) Expected | apparent MW: 103 kDa | 117 kDa
- Reactivity
-
Confirmed reactivity: Arabidopsis thaliana, Chlamydomonas reinardtii, red alga Gracilaria gracilis, Hordeum vulgare,Leptodictyum riparium (Hedw.) Warnst (moss), Medicago sativa, Phaeodactylum tricornutum Bohlin accession Pt1 8.6, Panax notoginseng, Populus yunanensis Dode, Solanum lycopersicum, Solanum tuberosum, Symbiodiniaceae of the genus Breviolum, Thalassiosira sp. (diatom), Trebouxia sp.,Vigna radiata, Vitis vinigera
Predicted reactivity: Arabis alpina, Brachypodium distachyon, Brassica napus, Brassica rapa subsp. pekinensis, Capsella rubella, Citrus clementina, Citrus sinensis, Chlorella vulgaris, Dunaliella salina, marine Diatoms, Coffea canephora, Eucalyptus grandis, Glycine max, Glycine soja, Gossypium arboretum, Helianthus annuus, Lycopersicum esculentum, Morus alba, Nannochloropsis gaditana, Nicotiana tabacum, Nicotiana attenuata, Nicotiana benthamiana, Oryza sativa, Phaseolus vulgaris, Phytophthora infestans, Physcomitrium patens, Prunus persica, Ricinus communis, Sorghum bicolor, Spinacia oleracea, Solanum lycopersicum, Symbiodinium microadriaticum, Theobroma cacao, Zea mays
Species of your interest not listed? Contact usNot reactive in: Aspergilus niger, Emiliania huxleyi, Tisochrysis lutea
- Application Examples
-
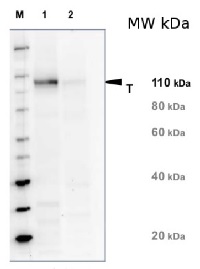
Application example 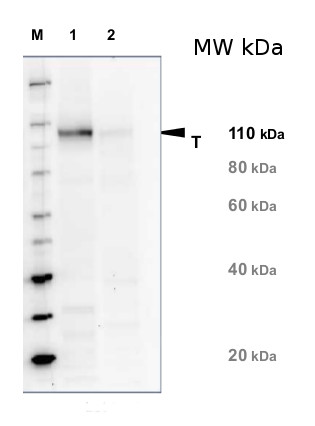
20 µg of total protein from Arabidopsis thaliana leaf (1) and Hordeum vulgare leaf (2) were extracted with Protein Extraction Buffer PEB (AS08 300). Samples were diluted with 1X sample buffer (NuPAGE LDS sample buffer (Invitrogen) supplemented with 50 mM DTT and heat at 70°C for 5 min and keept on ice before loading. Protein samples were separated on 4-12% Bolt Plus gels, LDS-PAGE and blotted for 70 minutes to PVDF using tank transfer. Blots were blocked immediately following transfer in 2% blocking reagent or 5% non-fat milk dissolved in 20 mM Tris, 137 mM sodium chloride pH 7.6 with 0.1% (v/v) Tween-20 (TBS-T) for 1h at room temperature with agitation. Blots were incubated in the primary antibody at a dilution of 1: 5 000 (in blocking reagent) for 1h at room temperature with agitation. The antibody solution was decanted and the blot was rinsed briefly twice, and then washed 1x15 min and 3x5 min with TBS-T at room temperature with agitation. Blots were incubated in secondary antibody (anti-rabbit IgG horse radish peroxidase conjugated, recommended secondary antibody AS09 602, Agrisera) diluted to 1:20 000 in blocking reagent for 1h at room temperature with agitation. The blots were washed as above. The blot was developed for 5 min with chemiluminescent detection reagent pf extreme femtogram range, according the manufacturers instructions. Images of the blots were obtained using a CCD imager (VersaDoc MP 4000) and Quantity One software (Bio-Rad). Exposure time was 5 minutes.Application examples: 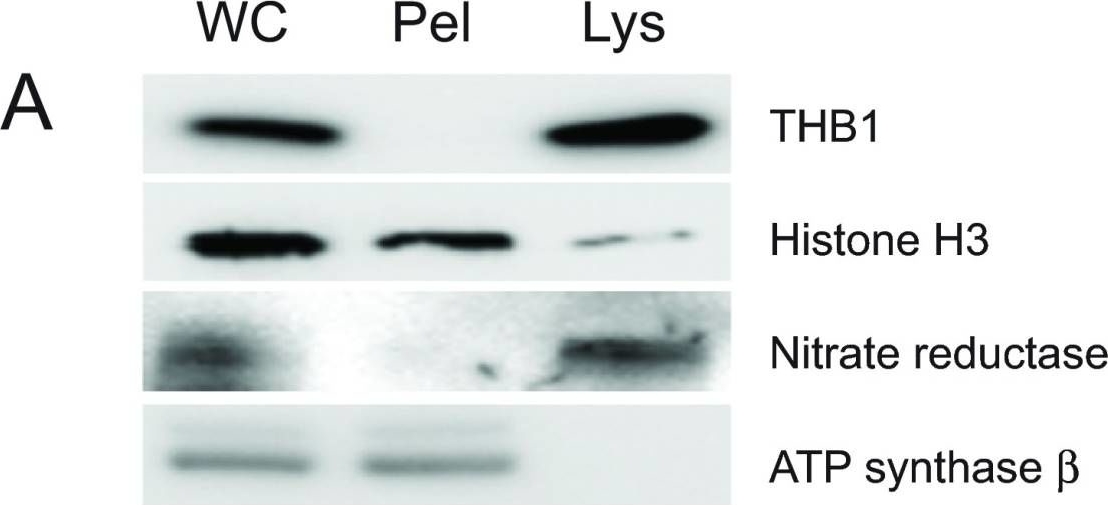
Reactant: Chlamydomonas reinhardtii (Green Alga)
Application: Western Blotting
Pudmed ID: 25653846
Journal: F1000Res
Figure Number: 1A
Published Date: 2015-02-06
First Author: Johnson, E. A. & Lecomte, J. T.
Impact Factor: None
Open PublicationPurification of THB1 fromC. reinhardtii cell culture.Protein samples from different stages of purification were analyzed by electrophoresis. (A) Samples of protein extracts before and after lysis ofC. reinhardtii cells with liquid nitrogen. Proteins separated on 16.5% Tris-tricine gel then transferred to nitrocellulose and immunostained with antibodies against proteins localized to different cellular compartments. Lysis by liquid nitrogen enriches the lysate with soluble proteins without enrichment of proteins found in major algal organelles. Histone H3 protein is located in the nucleus, nitrate reductase is a soluble cytosolic protein and the ATP synthase ? subunit is part of the thylakoid membrane of the chloroplast. (B) Samples from different steps in the purification procedure were separated by electrophoresis. Following separation, the proteins within the gel were visualized using silver stain. (C) Proteins prepared identically to those detected in panel B were transferred to nitrocellulose followed by immunostaining with polyclonal antibodies specific for THB1. In addition to the protein samples, a lane was used for molecular weight markers (Spectra LR, ThermoScientific). The numbers indicated between the panels represents location of the markers (kDa molecular mass). WC, whole cell protein extract. Lys, protein extract lysate, and Pel, protein pellet following liquid nitrogen fracturing. QFF, concentrated sample following anion exchange chromatography. S75, concentrated sample following separation on the Superdex 75 column.
Reactant: Chlorella
Application: Western Blotting
Pudmed ID: 33418923
Journal: Int J Mol Sci
Figure Number: 4A
Published Date: 2021-01-06
First Author: Kim, J., Chang, K. S., et al.
Impact Factor: 5.542
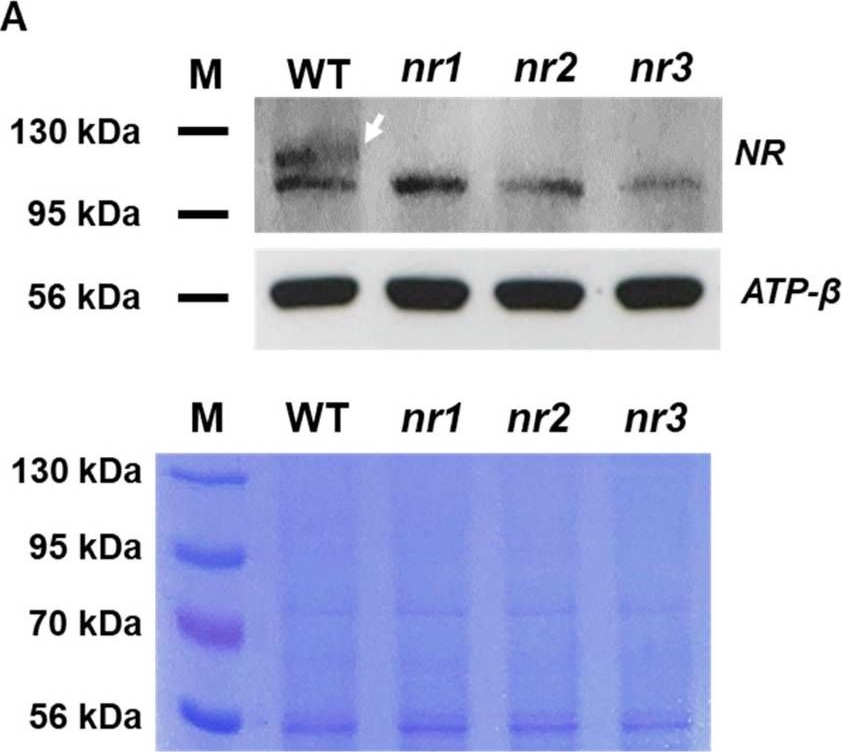
Open PublicationExpression of NR and APT in edited mutants compared with the WT. (A) Western blot analysis with a specific antibody against nitrate reductase (upper panel). Total protein content was assessed by Coomassie staining (lower panel). ATP-? was used as a reference protein. (B) Comparison of gene and amino acid sequence of APT between edited mutants and the WT. PCR amplification of APT from cDNA of apt mutants and WT cells (upper panel) and corresponding sequences (lower panels), showing the presence of frameshift mutations. M: size marker.
- Additional Information
-
Additional information (application): In Chlamydmonas reinhardtii anti-NR antibody is also reacting with L-Aminoacid Oxidase (a nitrogen scavenging enzyme induced during nitrogen starvation).
Using this antibody genome editing in Chlorella vulgaris UTEX395 by CRISPR-Cas9 system has been demonstrated as described in Kim et al. (2021)
Chemiluminescent detection is advised for NR detection using this antibody. - Background
-
Background: Assimilatory nitrate reductase (NR), (EC.1.6.6.1) catalyses the reduction of nitrate to nitrite in the cytoplasm. Plants contain 2 forms of NR: NADH-NR (most common form in plants and algae, predominantly found in green tissues) and NAD(P)H-NR (uses NADH or NADPH as the electron donor, constitutively expressed in plants at a low level). NADH-NR is a homodimer of two identical subunits (100-115 kDa each, hold together by a Mo-cofactor) each of them coded by up to three genes (NR1-3, NIA1-NIA3).
- Product Citations
-
Selected references: Expósito et al. (2023). Symbiotic interactions in the lichen R. farinacea dramatically modify NO biosynthetic source in Trebouxia microalgae.Algal Research Volume 75, September 2023, 103247.
Cao et al. (2022) Autophagic pathway contributes to low-nitrogen tolerance by optimizing nitrogen uptake and utilization in tomato. Hortic Res. 2022 Mar 23;9:uhac068. doi: 10.1093/hr/uhac068. PMID: 35669705; PMCID: PMC9164271.
Costa-Broseta et al. (2021). Post-Translational Modifications of Nitrate Reductases Autoregulates Nitric Oxide Biosynthesis in Arabidopsis. Int J Mol Sci. 2021 Jan 7;22(2):E549. doi: 10.3390/ijms22020549. PMID: 33430433.
Kim et al. (2021). Establishment of a Genome Editing Tool Using CRISPR-Cas9 in Chlorella vulgaris UTEX395. Int J Mol Sci. 2021 Jan 6;22(2):E480. doi: 10.3390/ijms22020480. PMID: 33418923.
Prinsi et al. (2021). Biochemical and Proteomic Changes in the Roots of M4 Grapevine Rootstock in Response to Nitrate Availability. Plants 10, no. 4: 792. https://doi.org/10.3390/plants10040792
Maresca et al. (2021) Biological responses to heavy metal stress in the moss Leptodictyum riparium (Hedw.) Warnst. Ecotoxicol Environ Saf. 2022 Jan 1;229:113078. doi: 10.1016/j.ecoenv.2021.113078. Epub 2021 Dec 17. PMID: 34929502.
Zhang et al. (2020). Hydrogen sulfide and rhizobia synergistically regulate nitrogen (N) assimilation and remobilization during N deficiency-induced senescence in soybean. Plant Cell Environ. 2020 Feb 3. doi: 10.1111/pce.13736.
Dongxu et al. (2020). Magnesium reduces cadmium accumulation by decreasing the nitrate reductase-mediated nitric oxide production in Panax notoginseng roots. Journal of Plant Physiology. Available online 7 February 2020, 153131.
Jayawardena et al. (2016). Elevated CO2 plus chronic warming reduces nitrogen uptake and levels or activities of nitrogen -uptake and -assimilatory proteins in tomato roots. Physiol Plant. 2016 Nov 28. doi: 10.1111/ppl.12532. [Epub ahead of print].
Chen et al. (2016). The role of nitric oxide signalling in response to salt stress in Chlamydomonas reinhardtii. Planta. 2016 Sep;244(3):651-69. doi: 10.1007/s00425-016-2528-0. Epub 2016 Apr 26.
Cheng et al. (2015). Quantitative proteomics analysis reveals that S-nitrosoglutathione reductase (GSNOR) and nitric oxide signaling enhance poplar defense against chilling stress. Planta. 2015 Aug 2.
Zhang et al. (2014). Heterologous expression of AtPAP2 in transgenic potato influences carbon metabolism and tuber development. FEBS Lett. 2014 Aug 27. pii: S0014-5793(14)00621-8. doi: 10.1016/j.febslet.2014.08.019.
Beyzaei et al. (2014). Response of Nitrate Reductase to Exogenous Application of 5-Aminolevulinic Acid in Barley Plants. J. Plant Growth Regulation, April 2014.
Frada et al. (2013). Quantum requirements for growth and fatty acid biosynthesis in the marine diatom Phaeodactylum tricornutum (Bacilloriophyceae) in nitrogen replete and limited conditions. J. Phycology. Diatom growth and lipid efficiency. - Protocols
-
Agrisera Western Blot protocol and video tutorials
Protocols to work with plant and algal protein extracts
Oxygenic photosynthesis poster by prof. Govindjee and Dr. Shevela
Z-scheme of photosynthetic electron transport by prof. Govindjee and Dr. Björn and Dr. Shevela - Reviews:
-
Ma?gorzata Reda | 2019-10-31We have been used this antibody for western blot detection of NR in cucumber (Cucumis sativus L.) roots tissue. 20 ug protein were separated by SDS-PAGE on 7.5% polyacrylamide gel, dilution of antibody was 1:2000. We observed clear product at about 100kDa. Obtain results were published in Reda et al. (2018) Involvement of NR and PM-NR in NO biosynthesis in cucumber plants subjected to salt stress, Plant Sci 267: 55-64