1
Anti-Xylose
AS07 267 | Clonality: Polyclonal | Host: Rabbit | Reactivity: Higher plants and algae
- Product Info
-
Immunogen: Xylose residues bound to the N-glycan in beta 1,2 Host: Rabbit Clonality: Polyclonal Purity: Immunogen affinity purified serum in PBS pH 7.4. Format: Lyophilized Quantity: 50 µg Reconstitution: For reconstitution add 50 µl of sterile water Storage: Store lyophilized/reconstituted at -20°C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Western blot (WB) Recommended dilution: 1 µg/ml (ELISA), 2 µg/10 ml incubation buffer (WB) Expected | apparent MW: 10-100 for various glycoproteins
- Reactivity
-
Confirmed reactivity: Higher plants and algae Predicted reactivity: Higher plants Not reactive in: No confirmed exceptions from predicted reactivity are currently known - Application Examples
-
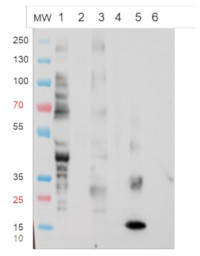
1 - 50 ug of cytosolic proteins from Arabidopsis thaliana WT
2 - 50 ug of cytosolic proteins from Arabidopsis thaliana fut11/fut12 mutant
3 - 50 ug of cytosolic proteins from Arabidopsis thaliana xylt mutant
4- 50 ug of cytosolic proteins from Arabidopsis thaliana xylt/ fut11/fut12 mutant
5- 1 ug of recombinant Avidin produced in maize (SIGMA)
6- 1 ug of Bovine Ribonuclease B (SIGMA) Mark: MW markers: PAGE Ruler pre-stained protein ladder from Fisher scientific
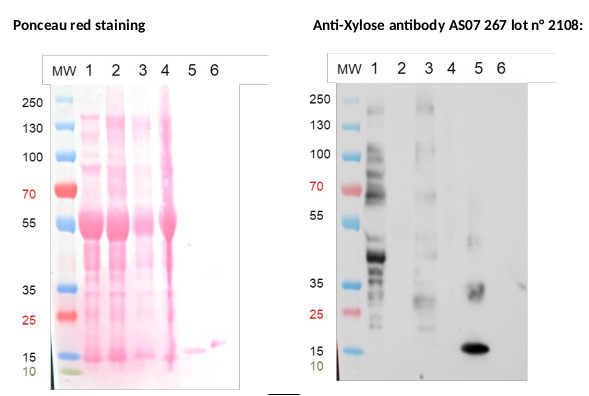
Protein extractions of about 500 mg of leaf of Arabidopsis thaliana in 1 mL Tris Buffer 0.1 M pH 7.5 were carried out in a high-speed benchtop homogenizer (FastPrep®-24 MP) 6 m.s-1, 40 s, 2 times, followed by a centrifugation step for 5 min at 10 000 rpm, 4°C. The pellet is discarded while the supernatant contains what we called cytosolic proteins. 20 µg of cytosolic proteins (WT and mutants Fut11/Fut12 ; xyltT ; xylt/fut11/Fut12) denaturated with leammli buffer at 100°C for 5 min were separated on Novex Bis-Tris 4-12% gel in MOPS buffer, and blotted on nitrocellulose using semi-dry transfer Thermo Scientific™ Pierce™ Power Blotter for 13 min in one step transfert buffer (thermoScientific 84731). Incubation of the blot is carried out with SNAP i.d.® 2.0 Protein Detection System according to the technical specifications of the supplier. Quickly, blot was blocked in TBST for 10 min, then the vacuum was applied. The primary antibody Anti-β(1,2)-Xylose (AS07 267) was used at a dilution of 1:3000 in TBST and left on the blot for 15 min before vacuum was applied. Three washes with TBST were performed and the secondary antibody (Goat anti rabbit IgG-HRP conjugated AS09 602) was used at a dilution 1:50 000 in TBST and left for 15 min on the blot before vacuum was applied. Three washes with TBST were performed and one last wash with TBS. The blot was developped for 3 min with Agrisera ECL set (AS16 ECL-SN-10) and time exposure is 30 sec performed with the VILBER Fusion FX Imager. We used Avidin (1µg) as positive control and Ribonuclease B (1µg) as negative control. Avidin carries β1,2 Xylose and ribonuclease B only mannose.
Conclusion: The Anti-(β1,2)-Xylose antibody (AS07 267) lot n° 2108 as expected, shows a positive response with Avidin, WT and fuT mutant and a negative response with the XylT mutant of Arabidopsis thaliana. It exhibits a negative response with ribonuclease B as expected too. Thus demonstrate the good specificity of the Anti-(β1,2)-Xylose antibody (AS07 267) lot n° 2108.
Courtesy of Dr Lehner and Mrs Burel, University of Rouen Normandie, FranceApplication example
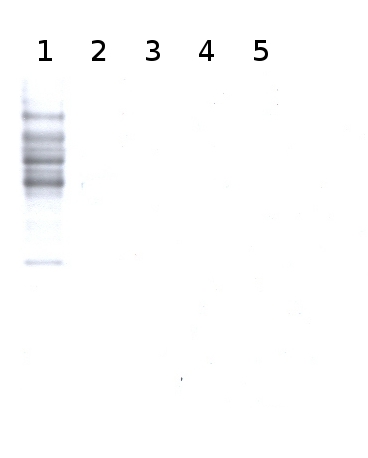
Total cell extract from Arabidopsis thaliana wild type (1) and cell extracts from different mutants defective in the complex N-glycan maturation pathway (2-5) (data not published yet).
Primary antibody has been used at 2 µg/10 ml of incubation buffer. Detection has been done using ECL.
Dot blot reaction of anti-Fucose and anti-Xylose antibodies with various controls: Avidin (Fuc+/Xyl+), Fetuin (Fuc-/Xyl-), PLA2 (Fuc+/Xyl-) and Mur1-2 (Fuc-/Xyl+). 2 µl of each extract were spotted on a nitrocellulose membrane, placed on top of 2 WHATMAN filters (one soaked in TBS-T) and dried for 1.5 h at RT. The mem-brane was blocked for 30 min with 2% low-fat milk powder in TBS-T (0.1% TWEEN 20) and incubated with anti-Fucose(1) (AS07 268, 1:1000) or anti-Xylose(2) (AS07 267, 1:1000) for 30 min and then with secondary anti-rabbit(1:1000) antibody (ALP conjugated, recommended secondary antibody AS09 607). Membrane was washed with TBS-T 3 x 10 minutes before reaction development using alkaline phosphatase reagent BCIP®/NTB premixed solution (Sigma, Prod. No. B6404).
Please follow this link for a more detailed Dot-Blot protocol
- Additional Information
-
Additional information: Beta (1,2) xylose is present exclusively in plant N-glycans so antibodies against this sugar moiety should not cross-react with any mammal glycoprotein.
This antibody do not bind free D-xylose. This antibody does not seem to work in immunolocalization.Additional information (application): Negative control: Fetuin, a glycoprotein containing fucose linked in alpha 1.6 and no xylose, Sigma, product number F3385.
Positive control: Type II - horseradish peroxidase which contains β1.2 Xylose and α1.3 fucose, Sigma, product number P8250
- Background
-
Background: This antibody specifically cross-reacts against xylose residues bound to the protein N-glycans in beta1,2. This residue is characterisitc of the plant protein N-glycans and is absent in protein N-glycans from animals. This residue is added in the Golgi apparatus.
- Product Citations
-
Selected references: Mazalovska and Kouokam (2026). Transiently Expressed Mistletoe Lectin II in Nicotiana benthamiana Demonstrates Anticancer Activity In Vitro. Molecules. 2020 May 31;25(11):2562. doi: 10.3390/molecules25112562.
Petrova et al. (2026). Recombinant Flax Lectin from Legume Family: Structure Modeling and Analysis of Posttranslational Modifications. Dokl Biochem Biophys. 2025 Dec;525(2):632-638. doi: 10.1134/S1607672925601453.
Xu et al. (2023). The Golgi-localized transporter OsPML4 contributes to manganese homeostasis in rice. Plant Sci. 2023 Dec 2:339:111935.doi: 10.1016/j.plantsci.2023.111935.
Ropitaux et al. (2023). Subcellular localization of core beta (1, 2)-xylosylated N-glycoproteins in the green microalgae Chlamydomonas reinhardtii. Algal Research Volume 77, January 2024, 103366.
Xavier et al. (2021) Inactivation of N-Acetylglucosaminyltransferase I and a1,3-Fucosyltransferase Genes in Nicotiana tabacum BY-2 Cells Results in Glycoproteins With Highly Homogeneous, High-Mannose N-Glycans. Frontiers in Plant Science. Volume 12. doi: 10.3389/fpls.2021.634023
Yang et al. (2021). Golgi-localized manganese transporter PML3 regulates Arabidopsis growth through modulating Golgi glycosylation and cell wall biosynthesis. New Phytol. 2021 Jan 17. doi: 10.1111/nph.17209. Epub ahead of print. PMID: 33454966.
Lucas et al. (2019). Multiple xylosyltransferases heterogeneously xylosylate protein N-linked glycans in Chlamydomonas reinhardtii. Plant J. 2019 Nov 27. doi: 10.1111/tpj.14620.
Lucas et al. (2019). Multiple xylosyltransferases heterogeneously xylosylate protein N-linked glycans in Chlamydomonas reinhardtii. Plant J. 2019 Nov 27. doi: 10.1111/tpj.14620.
Hou et al. (2019). Identification and characterization of a novel glycoprotein core xylosidase from the bacterium Elizabethkingia meningoseptica. Biochemical and Biophysical Research Communications Available online 27 July 2019.
Jansing et al. (2018). CRISPR/Cas9-mediated knockout of six glycosyltransferase genes in Nicotiana benthamiana for the production of recombinant proteins lacking Beta-1,2-xylose and core alfa-1,3-fucose. Plant Biotechnol J. 2018 Jul 3. doi: 10.1111/pbi.12981.
Nakanishi et al. (2017). Protection of Human Colon Cells from Shiga Toxin by Plant-based Recombinant Secretory IgA. Sci Rep. 2017 Apr 3;7:45843. doi: 10.1038/srep45843. (ELISA).
Hanania et al. (2017). Establishment of a tobacco BY2 cell line devoid of plant specific xylose and fucose as a platform for the production of biotherapeutic proteins. Plant Biotechnol J. 2017 Feb 3. doi: 10.1111/pbi.12702.
Ebert et al. (2015). Identification and Characterization of a Golgi-Localized UDP-Xylose Transporter Family from Arabidopsis. Plant Cell. 2015 Mar 24. pii: tpc.114.133827.
Mathieu-Rivet et al. (2013). Exploring the N-glycosylation pathway in Chlamydomonas reinhardtii unravels novel complex structures. Mol Cell Proteomics, Aug 2. - Protocols
-
Agrisera Western Blot protocol and video tutorials
Protocols to work with plant and algal protein extracts - Reviews:
-
This product doesn't have any reviews.