1
Anti-PEPC | Phosphoenolpyruvate carboxylase
AS09 458 | Clonality: Polyclonal | Host: Rabbit | Reactivity: A. comosus, A. thaliana, C. ciliaris, C. gayana, C. velia, Ch. quinoa, H. vulgare, J. curcas, K. prostrata, L. fusca, Lupinus sp. , M. maximus, M. crystallinum, N. tabacum, O. sativa, P. antidotale, P. coloratum, P. strobus, Saccharum spp. hybrid clone C91-301, S. lanata, S. laricifolia, S. bicolor, Synechocystis PCC 6803, Phaeodactylum tricornutum (strain CCAP 1055/1), T. weissfloggi, Z. mays, Z. muelleri
Benefits of using this antibody
- Product Info
-
Immunogen: KLH-conjugated synthetic peptide well conserved PEPC1 and sequences from different plant species including Arabidopsis thaliana Q9MAH0, At1g53310 (PEPC 1), Q84VW9, At3g14940 (PEPC 3). The peptide chosen to elicit this antibody is also perfectly conserved in bacterial type of this enzyme NP_177043.2 (PEPC 4).
For Zea mays, the peptide is converved in PEP1 and PEP4 isoforms.Host: Rabbit Clonality: Polyclonal Purity: Immunogen affinity purified serum in PBS pH 7.4. Format: Lyophilized Quantity: 50 µg Reconstitution: For reconstitution add 50 µl of sterile water Storage: Store lyophilized/reconstituted at -20°C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Please do not re-use this primary antibody solution. In case of cyanobacterial samples there will be no signal in your second incubation. Tested applications: Immunolocalization (IL), Western blot (WB) Recommended dilution: 1 : 500 (IL), 1: 1000 - : 10 000 (WB) Expected | apparent MW: 110 | 105 kDa - Reactivity
-
Confirmed reactivity: Ananas comosus, Arabidopsis thaliana, Cenchrus ciliaris, Chenopodium quinoa, Chloris gayana, Chromera velia, Cyanthobasis fruticulosa, Glycine max, Hordeum vulgare, Jatropha curcas, Kochia prostrata, Leptochloa fusca, Lupinus sp. , Megathyrsus maximus, Mesembryanthemum crystallinum, Nicotiana tabacum, Oryza sativa, Panicum antidotale, Panicum coloratum, Petrosimonia nigdeensis, Pinus strobus, Saccharum spp. hybrid clone C91-301, Salsola lanata, Salsola laricifolia,Salsola grandis, Salsola tragus Sorghum bicolor, Synechocystis PCC 6803, Phaeodactylum tricornutum (strain CCAP 1055/1), Pinus strobus, Thalassiosira weissfloggi, Zea mays, Zostera muelleri
Predicted reactivity: Amaranthus sp., Brassica napus, Cucumis sativus (PEPC1, PEPC2, PEPC3), Flaveria bidentis, Flaveria trinervia, Lupinus albus, Mammillaria thornberi, Manihot esculenta, Manihot obovata, Medicago sativa, Morinda citrifolia, Nannochloropsis gaditana CCMP526, Nopalea gaumeri, Opuntia macbridei, Pachycereus pringlei, Pachycereus hollianus, Pisum sativa, Phaseolus vulgaris, Phragmites australis, Populus sp.,Saccharum spp, Solanum tuberosum, Spinacia oleracea, Streptanthus tortuosus,Triticum aestivum, algae, diatoms: Thalassiosira pseudonana, other species: Salmonella sp., Schiedea hookeri, Shigella sp. Schiedea sarmentosa, Solanum lycopersicum, Streptanthus farnsworthianus, Tacinga saxatilis,Yersinia sp. Vibrio sp., Quercus sp.
Species of your interest not listed? Contact usNot reactive in: Methanothermobacter thermautotrophicus - Application Examples
-
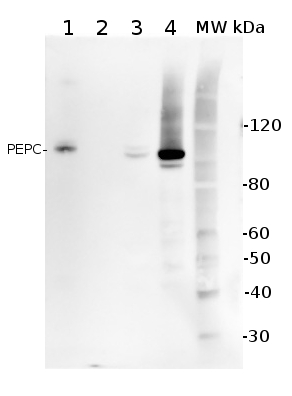
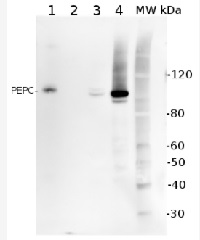
5 µg of total protein from (1) Arabidopsis thaliana leaf extracted with Protein Extration Buffer, PEB (AS08 300), (2) Spinacia oleracea total cell, extracted with PEB, (3) Hordeum vulgare total cell extracted with PEB, (4) Zea mays total cell extracted with PEB, were separated on 4-12% NuPage (Invitrogen) LDS-PAGE and blotted 1h to PVDF. Blots were blocked immediately following transfer in 2% blocking reagent (GE Healthcare) in 20 mM Tris, 137 mM sodium chloride pH 7.6 with 0.1% (v/v) Tween-20 (TBS-T) for 1h at room temperature with agitation. Blots were incubated in the primary antibody at a dilution of 1: 10 000 for 1h at room temperature with agitation. The antibody solution was decanted and the blot was rinsed briefly twice, then washed once for 15 min and 3 times for 5 min in TBS-T at room temperature with agitation. Blots were incubated in secondary antibody (anti-rabbit IgG horse radish peroxidase conjugated) diluted to 1:50 000 in 2% blocking solution for 1h at room temperature with agitation. The blots were washed as above and developed for 5 min with chemiluminescent detection reagent according the manufacturers instructions. Images of the blots were obtained using a CCD imager (FluorSMax, Bio-Rad) and Quantity One software (Bio-Rad).
PEPC is especially prone to proteolysis. To avoid artifacts, we use chymotrypsin, as recommended by Plaxton (2019), in addition to the Roche protease inhibitor cocktail.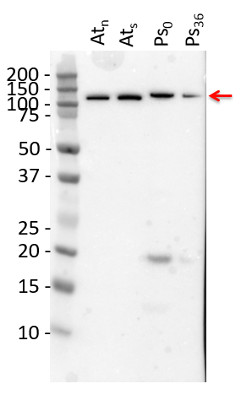
10 µg of total protein extracted freshly from Arabidopsis thaliana wt leaf tissue (Atn non-senescent leaves), Arabidopsis thaliana wt leaf tissue (Ats senescent leaves), Pinus strobus needle tissue (PS0, PS36 ) with 1 M Tris-HCl, pH 6.8, 10 % SDS, 15 % sucrose, 0.5 DTT and denatured at 75°C for 5 min. were separated on 10 % Bis-Tris Nupage Novex gel (120 V/45 min. using MES buffer system) and blotted 30 min. to PVDF. Blot was blocked with 5 % non-fat milk 45 min./RT with agitation. Blot was incubated in the primary antibody at a dilution of 1: 1000 for 1h/RT with agitation in TBS with 2 % non-fat milk or ON/4°C with agitation. The antibody solution was decanted and the blot was rinsed briefly twice for 10 min. in TBS at RT with agitation. Blot was incubated in Agrisera matching secondary antibody (anti-rabbit IgG horse radish peroxidase conjugated, AS09 602) diluted to 1:75 000 in for 1h/RT with agitation. The blot was washed as above and developed using chemiluminescent detection. Exposure time was 40 seconds.
Courtesy of Dr. Christine Yao-Yun Chang and the Ensminger lab, University of Toronto, CanadaApplication examples: 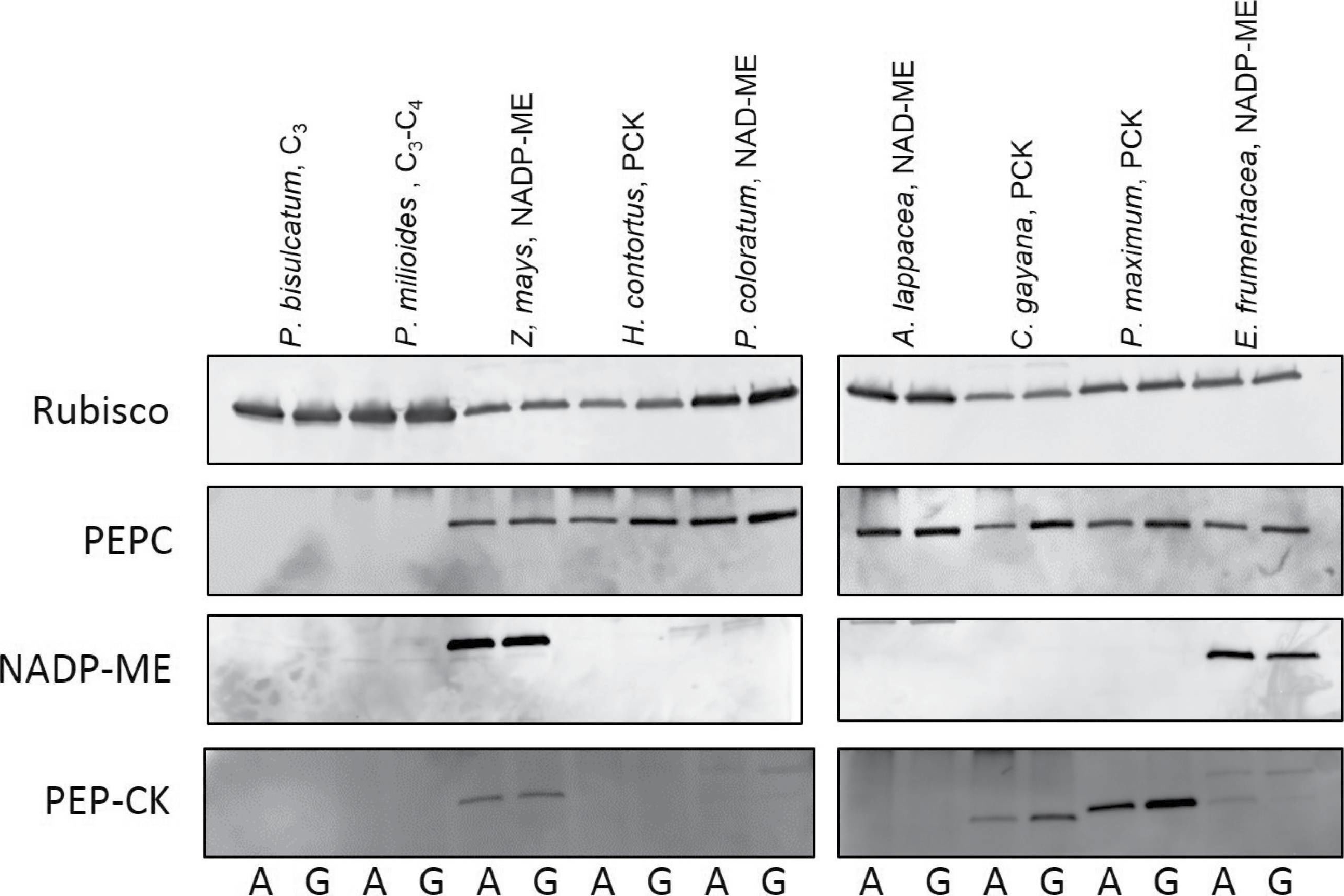
Reactant: Arabidopsis thaliana (Thale cress)
Application: Western Blotting
Pudmed ID: 24723409
Journal: J Exp Bot
Figure Number: 4A
Published Date: 2014-07-01
First Author: Pinto, H., Sharwood, R. E., et al.
Impact Factor: 6.088
Open PublicationImmunoblot analyses of photosynthetic enzymes. Examples of immunoblot analysis for the photosynthetic proteins Rubisco (A), PEPC (B), NADP-ME (C), and PEP-CK (D) extracted from leaves of selected grass species grown at glacial (180 ?l l–1, G) or ambient (400 ?l l–1, A) [CO2].

Reactant: Zea mays (Maize/Corn)
Application: Western Blotting
Pudmed ID: 26208645
Journal: J Exp Bot
Figure Number: 10C
Published Date: 2015-11-01
First Author: Chen, J., Wu, F. H., et al.
Impact Factor: 6.088
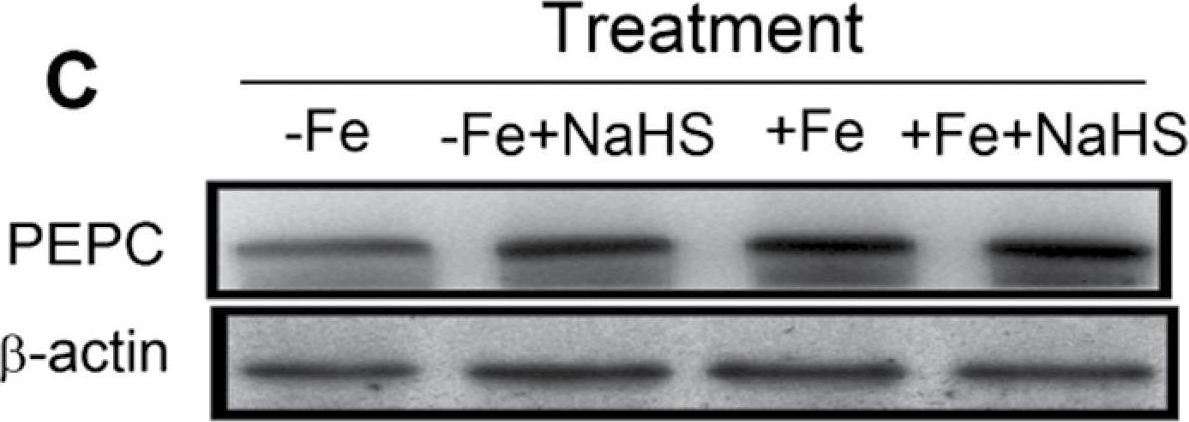
Open PublicationWestern blot analysis of RuBISCO LSU (A) and PEPC (C) of maize plants. Maize seedlings were pre-treated with 100 µM NaHS for 8 d and then grown in a nutrient solution containing 1 µM Fe(III)-EDTA or 50 µM Fe(III)-EDTA for 12 d. Relative expression level is shown as the ratio of RuBISCO LSU:?-actin (B) and PEPC:?-actin (D) using Quantity One software. Data are presented as means ± SE. Columns labelled with different letters indicate significant differences at P<0.05. –Fe, 1 µM Fe; –Fe+NaHS, seedlings were pre-treated with 100 µM NaHS and then treated with 1 µM Fe; +Fe, 50 µM Fe; +Fe+NaHS, seedlings were pre-treated with 100 µM NaHS and then treated with 50 µM Fe.
Reactant: Echinochloa (Barnyard grass)
Application: Western Blotting
Pudmed ID: 29659931
Journal: J Exp Bot
Figure Number: 5A
Published Date: 2018-05-25
First Author: Sonawane, B. V., Sharwood, R. E., et al.
Impact Factor: 6.088
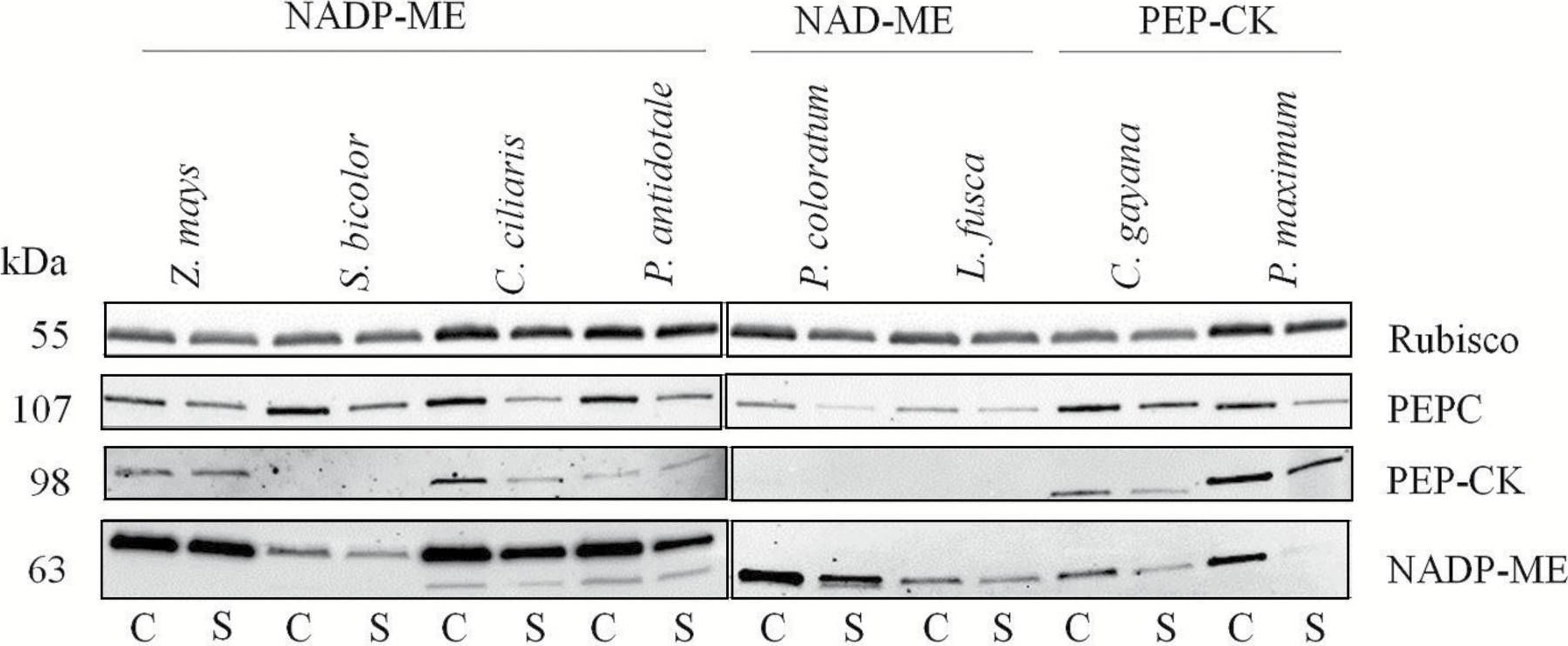
Open PublicationImmunoblot analysis of photosynthetic enzymes. Immunoblot analysis for the photosynthetic proteins Rubisco, PEPC, PEP-CK, and NADP-ME extracted from leaves of eight C4 grasses belonging to three biochemical subtypes in control (C) or shade (S) environments. Loaded volumes varied between 4 ?l and 15 ?l in order to normalize the protein content to a common leaf area. Because of the small gel size, a limited number of samples (8–9) was loaded on an individual gel. Finally, all immunoblots of the studied protein and species were arranged in a composite figure. For uniform visualization, gamma settings of individual images were adjusted. A protein ladder was used for individual immunoblots; for simplicity, band size is referred to numerically.
Reactant: Oryza sativa (Asian rice)
Application: Western Blotting
Pudmed ID: 30499169
Journal: Mol Plant Pathol
Figure Number: 3A
Published Date: 2019-04-01
First Author: Hui, S., Shi, Y., et al.
Impact Factor: 5.418
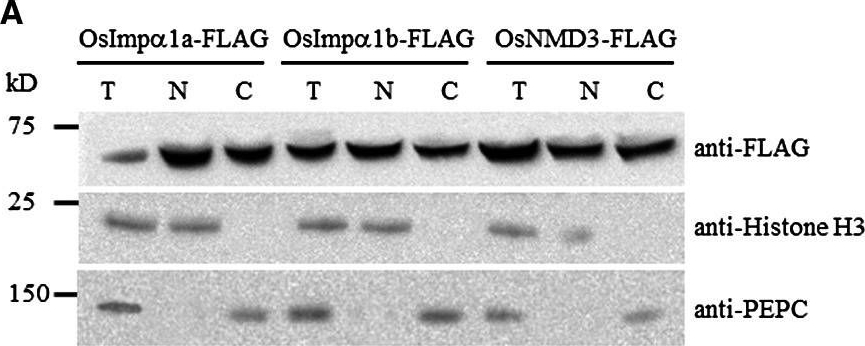
Open PublicationExpression patterns of OsImp?1a and OsImp?1b. (A) OsImp?1a and OsImp?1b are nucleocytoplasmic?localized proteins. Total protein, the nucleus?depleted fraction and nucleus?enriched fraction were loaded onto a sodium dodecylsulfate?polyacrylamide gel electrophoresis (SDS?PAGE) gel and subjected to immunoblot analysis. Histone H3 and phosphoenolpyruvate carboxylase (PEPC) were used as nuclear and cytosolic markers, respectively. T, total protein extracts; N, nucleus?enriched fraction; C, nucleus?depleted fraction. (B) Expression of OsImp?1a and OsImp?1b in different tissues. Tissues were collected at the booting stage from IR24. (C) Expression of OsImp?1a and OsImp?1b after infection with Xanthomonas oryzae pv. oryzae (Xoo) strain PXO99.
- Additional Information
-
Additional information: PEPC is especially prone to proteolysis. To avoid artifacts, we use chymotrypsin, as recommended by Plaxton (2019), in addition to the Roche protease inhibitor cocktail. Additional information (application): Antibody can be also used following 2D gel electrophoresis - Background
-
Background: PEPC (phosphoenolpyruvate carboxylase), EC=4.1.1.31, belongs to an enzyme family of carboxy-lyases that is catalyzing adding fo carbon dioxide to phosphoenolpyruvate (PEP) to form oxaloacetate. Alternative names: PEPCase 1, PEPCase 3, PEPC 1, PEPC 3
- Product Citations
-
Selected references: Nosek et al. (2026). Can continuous light mitigate the detrimental actions of salinity stress in facultative CAM plants? Photosynth Res . 2026 Mar 23;164(2):23. doi: 10.1007/s11120-026-01211-7.
Rakhmankulova et al. (2026). Effect of drought and elevated temperature on the physiological and biochemical properties of C3 and C4 halophytes in Amaranthaceae. Journal of Arid Land Volume 18, Issue 1, January 2026, Pages 131-149.
Gong et al. (2025). Far-red light orchestrates antiviral defense in plants. Sci Adv. 2025 Nov 21;11(47):eadz7663. doi: 10.1126/sciadv.adz7663.
Cruz et al. (2025). Canopy-Level Photosynthetic Homogeneity Accounts for Greater Biomass in Energy Cane Compared to Sugarcane. Tropical Plant Biol. 18, 81 (2025). doi.org/10.1007/s12042-025-09451-y.
Kocacinar et aL. (2025). Biochemical and structural differences between C3 cotyledons and C4 leaves in species of Salsoloideae (Chenopodiaceae). Sci Rep. 2025 Oct 17;15(1):36383.doi: 10.1038/s41598-025-20388-w.
Nosek et al. (2025). Differential expression of RubisCO and PEPC during the salinity stress recovery of facultative CAM plants. Plant Physiol Biochem. 2025 Jul 18:228:110272. doi: 10.1016/j.plaphy.2025.110272
Fang et al. (2024). Subfunctionalisation and self-repression of duplicated E1 homologues finetunes soybean flowering and adaptation. Nat Commun. 2024 Jul 23;15(1):6184. doi: 10.1038/s41467-024-50623-3.
Nguyen et al.(2024). The processed C-terminus of AvrRps4 effector suppresses plant immunity via targeting multiple WRKYs.J Integr Plant Biol. 2024 Jun 13.doi: 10.1111/jipb.13710.
Cruz et al.(2023). Variation of photosynthesis along the canopy profile of sugarcane and energy canes. Research Square; DOI: 10.21203/rs.3.rs-3124093/v1
Luo, et al. (2023) Deubiquitinating enzymes UBP12 and UBP13 stabilize the brassinosteroid receptor BRI1. EMBO Rep. 2022;23(4):e53354. doi:10.15252/embr.202153355
Ding et al. (2022) CPK28-NLP7 module integrates cold-induced Ca2+ signal and transcriptional reprogramming in Arabidopsis. Sci Adv. 2022 Jul;8(26):eabn7901. doi: 10.1126/sciadv.abn7901. Epub 2022 Jun 29. PMID: 35767615; PMCID: PMC9242591.
Lan, Ma, Zheng, et al. (2022) Ubiquitome profiling reveals a regulatory pattern of UPL3 with UBP12 on metabolic-leaf senescence. Life Sci Alliance. 2022;5(12):e202201492. Published 2022 Aug 4. doi:10.26508/lsa.202201492
Durall et al. (2021). Production of succinate by engineered strains of Synechocystis PCC 6803 overexpressing phosphoenolpyruvate carboxylase and a glyoxylate shunt. Microb Cell Fact. 2021 Feb 8;20(1):39. doi: 10.1186/s12934-021-01529-y. PMID: 33557832; PMCID: PMC7871529.
Wang et al. (2021). Brassinosteroids inhibit miRNA-mediated translational repression by decreasing AGO1 on the endoplasmic reticulum. J Integr Plant Biol. 2021 May 21. doi: 10.1111/jipb.13139. Epub ahead of print. PMID: 34020507.
Rakhmankulova et al. (2021) Possible Activation of ?3 Photosynthesis in ?4 Halophyte Kochia prostrata Exposed to an Elevated Concentration of ??2. Russ J Plant Physiol 68, 1107–1114 (2021). https://doi.org/10.1134/S1021443721060169
Durall et al. (2020). Increased ethylene production by overexpressing phosphoenolpyruvate carboxylase in the cyanobacterium Synechocystis PCC 6803. Biotechnol Biofuels. 2020 Jan 28;13:16. doi: 10.1186/s13068-020-1653-y.
Kramer et al. (2020). N6?methyladenosine and RNA secondary structure affect transcript stability and protein abundance during systemic salt stress in Arabidopsis. Plant Direct . 2020 Jul 24;4(7):e00239.doi: 10.1002/pld3.239.
Wang et al. (2019). PUB25 and PUB26 Promote Plant Freezing Tolerance by Degrading the Cold Signaling Negative Regulator MYB15.
Hui et al. (2018). TALE-carrying bacterial pathogens trap host nuclear import receptors for facilitation of infection of rice. Mol Plant Pathol. 2018 Nov 30. doi: 10.1111/mpp.12772.
Salesse-Smith et al. (2018). Overexpression of Rubisco subunits with RAF1 increases Rubisco content in maize. Nat Plants. 2018 Oct;4(10):802-810. doi: 10.1038/s41477-018-0252-4.
Bassi et al. (2018). Nitrogen supply influences photosynthesis establishment along the sugarcane leaf. Sci Rep. 2018 Feb 2;8(1):2327. doi: 10.1038/s41598-018-20653-1.
Sonawane et al. (2018). Shade compromises the photosynthetic efficiency of NADP-ME less than PEP-CK and NAD-ME C 4 grasses. J Exp. Botany, doi.org/10.1093/jxb/ery129.
Wen et al. (2017). Possible involvement of phosphoenolpyruvate carboxylase and NAD-malic enzyme in response to drought stress. A case study: A succulent nature of the C4-NAD-ME type desert plant, Salsola lanata (Chenopodiaceae). Functional Plant Biology 44(12), DOI10.1071/FP16430
Jiang et al. (2017). Development of an Efficient Protein Extraction Method Compatible with LC-MS/MS for Proteome Mapping in Two Australian Seagrasses Zostera muelleri and Posidonia australis. Frontiers in Plant Science, doi: 10.3389/fpls.2017.01416.
Liu et al. (2017). Plasma Membrane CRPK1-Mediated Phosphorylation of 14-3-3 Proteins Induces Their Nuclear Import to Fine-Tune CBF Signaling during Cold Response. Mol Cell. 2017 Apr 6;66(1):117-128.e5. doi: 10.1016/j.molcel.2017.02.016.
Ribeiro et al. (2017). Increased sink strength offsets the inhibitory effect of sucrose on sugarcane photosynthesis. J Plant Physiol. 2017 Jan;208:61-69. doi: 10.1016/j.jplph.2016.11.005.
Shen et al. (2016). The existence of C4-bundle-sheath-like photosynthesis in the mid-vein of C3 rice. Rice (N Y). 2016 Dec;9(1):20. doi: 10.1186/s12284-016-0094-5. Epub 2016 May 10.
Ishikawa et al. (2016). NDH-Mediated Cyclic Electron Flow Around Photosystem I is Crucial for C4 Photosynthesis. Plant Cell Physiol. 2016 Aug 6. pii: pcw127. [Epub ahead of print]
Shen et al. (2015). Overexpression of maize phosphoenolpyruvate carboxylase improves drought tolerance in rice by stabilization the function and structure of thylakoid membrane. Photosynthetica, September 2015, Volume 53, Issue 3, pp 436-446.
Foley et. al (2015). Analysis of conglutin seed storage proteins across lupin species using transcriptomic, protein and comparative genomic approaches. BMC Plant Biology 2015, 15:106 doi:10.1186/s12870-015-0485-6. - Protocols
-
Agrisera Western Blot protocol and video tutorials
Protocols to work with plant and algal protein extracts
Oxygenic photosynthesis poster by prof. Govindjee and Dr. Shevela
Z-scheme of photosynthetic electron transport by prof. Govindjee and Dr. Björn and Dr. Shevela - Reviews:
-
Piotr Rozp?dek | 2014-03-24I confirm reactivity against Mesembryanthemum crystallinum. Dillution: as in instructionDaniela Ewe | 2011-11-07I can confirm reactivity against protein from Phaeodactylum tricornutum. The band is clear and there are no other disturbing bands at the same hight. Unfortunately, the antibody is not very stable so I can’t recommend recycling. I don’t use it for more than 3 days. Another aspect is the low dilution factor of 1:2000. This consumes a lot of antibody.
Related products
AS07 241 | Clonality: Polyclonal | Host: Rabbit | Reactivity: [global antibody] for plants Ananas comosus, Miscantus giganteus, Mouse, Nannochloropsis oceanica, Oryza sativa, Panicum maximum,, Panicum virgatum, Phaseolus vulgaris, Saccharum spp. hybrid clone C91-301, Spartina alterniflora, Spartina patens, Zea mays
Benefits of using this antibody