1
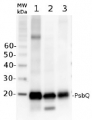
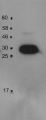
Anti-PsbO2 | 33 kDa of the oxygen evolving complex (OEC) of PSII
AS14 2825 | Clonality: Polyclonal | Host: Rabbit | Reactivity: A. thaliana
- Product Info
-
Immunogen: KLH-conjugated peptide chosen chosen from Arabidopsis thaliana PsbO2, UniProt:Q9S841, TAIR: At3g50820
Host: Rabbit Clonality: Polyclonal Purity: Serum Format: Lyophilized Quantity: 50 µl Reconstitution: For reconstitution add 50 µl of sterile water Storage: Store lyophilized/reconstituted at -20°C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Immunoprecipitation (IP), Western blot (WB) Recommended dilution: 1 : 1000 (WB) Expected | apparent MW: 33 | 30 kDa
- Reactivity
-
Confirmed reactivity: Arabidopsis thaliana
Predicted reactivity: Brassica napus, Capsella rubella, dinoflagellate, Triticum aestivum
Species of your interest not listed? Contact us - Application Examples
-
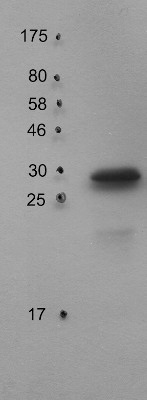
application example 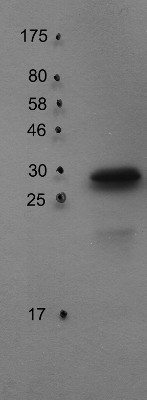
30 µg of thylakoid protein from Arabidopsis thaliana Columbia-0 extracted with 330 mM sorbitol, 50 mM Hepes-KOH (pH 7.8) were separated on 15% SDS-PAGE and blotted 1h to PVDF using semi-dry transfer. Blots were blocked with 5% milk for 1h at room temperature (RT) with agitation. Blot was incubated in the primary antibody at a dilution of 1: 1 000 for 6h at 4°C with agitation. The antibody solution was decanted and the blot was rinsed briefly twice, then washed once for 15 minutes and 3 times for 5 min in TBS-T at RT with agitation. Blot was incubated in secondary antibody (anti-rabbit IgG horse radish peroxidase conjugated) diluted to 1:10 000 in 1% milk (TBS-T) for 2h at 4°C with agitation. The blot was washed as above and developed for 5 min with ECL according to the manufacturer's instructions. Exposure time was 60 seconds.
Courtesy of Dr. Peter Gollan, University of Turku, FinlandApplication examples: 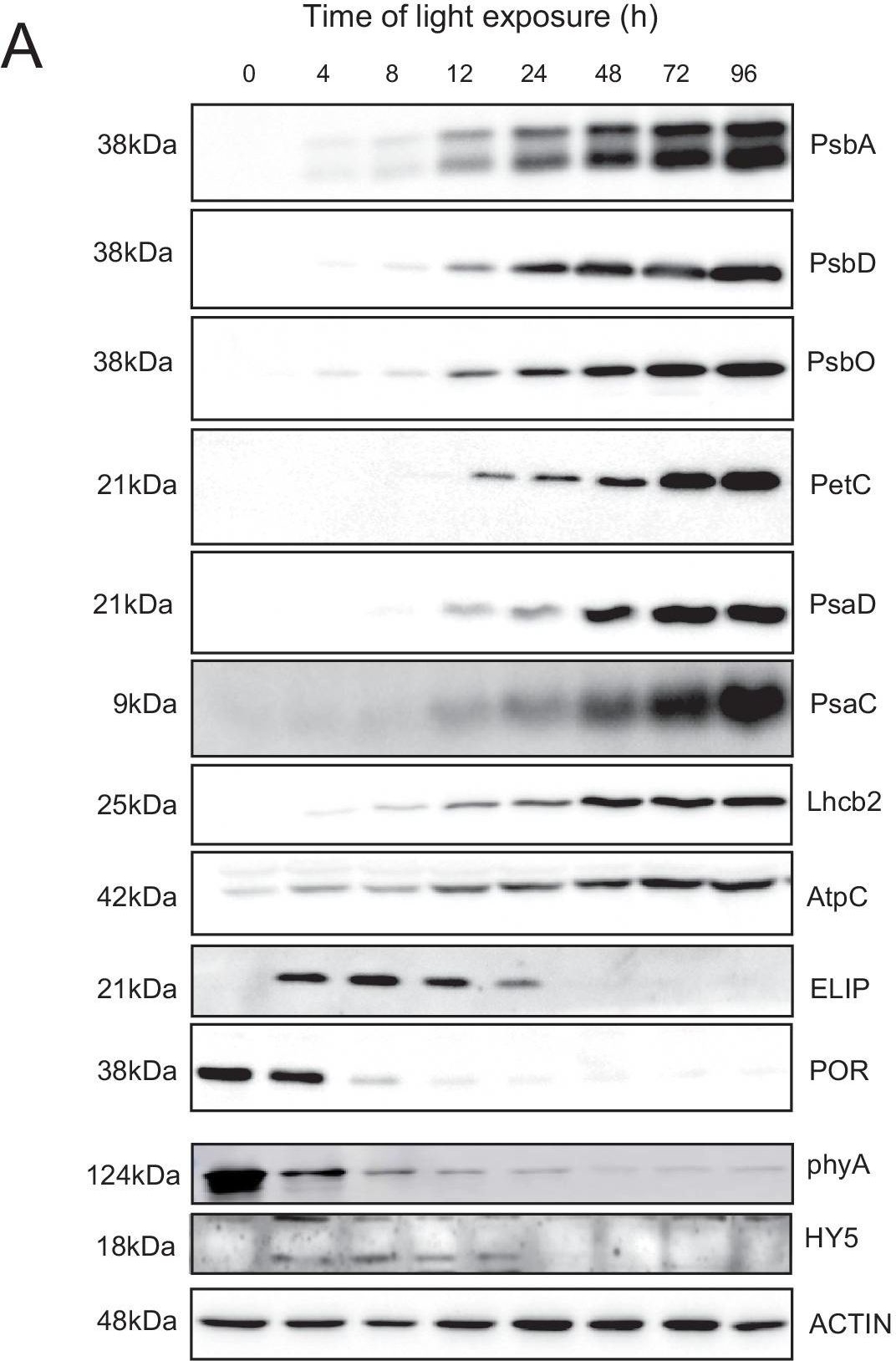
Reactant: Arabidopsis thaliana (Thale cress)
Application: Western Blotting
Pudmed ID: 33629953
Journal: Elife
Figure Number: 6A
Published Date: 2021-02-25
First Author: Pipitone, R., Eicke, S., et al.
Impact Factor: 7.448
Open PublicationAccumulation dynamics of photosynthesis-related proteins during de-etiolation.Three-day-old etiolated seedlings of Arabidopsis thaliana were illuminated for 0 hr (T0), 4 hr (T4), 8 hr (T8), 12 hr (T12), 24 hr (T24), 48 hr (T48), 72 hr (T72), and 96 hr (T96) under white light (40 µmol/m2/s). (A) Proteins were separated by SDS-PAGE and transferred onto nitrocellulose membrane and immunodetected with antibodies against PsbA, PsbD, PsbO, PetC, PsaD, PsaC, Lhcb2, AtpC, ELIP, POR, phyA, HY5, and ACTIN proteins. (B–C) Quantification of PsbA, PetC, and PsaC during de-etiolation. Heatmap (B) was generated after normalization of the amount of each protein relative to the last time point (T96). Graph (C) corresponds to the absolute quantification of proteins at T96. Error bars indicate ± SD (n = 3). Quantification of photosystem-related proteins during de-etiolation is detailed in Figure 6—figure supplement 1.Figure 6—source data 1.Quantitative data for immunoblot analysis.Quantitative data for immunoblot analysis.Quantification of photosynthesis-related proteins.(A) Immunodetection of PsbA, PetC, and PsaC during de-etiolation. Dilutions were used for the later time points to avoid saturation of the signal. (B) Different bands were detected by Amersham Imager program and quantified by Image QuantTL (Amersham). (C) Calibration curves were created using recombinant proteins (Agrisera). Calibration curve composition: PsbA 10 ng (A; lane a), 5 ng (b), 2.5 ng (c), and 1.25 ng (d); PetC 10 ng (e), 5 ng (f), 2.5 ng (g), and 1.25 ng (h); PsaC 3 ng (i), 1.5 ng (l), 0.75 ng (m), and 0.325 ng (n). Data indicate mean ± SD (n = 3–4). Raw data and calculations are shown in Figure 6—source data 1.
- Additional Information
-
Additional information (application): This antibody is specific to PsbO2 and does not react with PsbO1 as confirmed on deletion mutant - Background
-
Background: The PsbO protein is an extrinisic subunit of the water splitting photosystem II (PSII) complex. The protein is exposed on the luminal side of the thylakoid membrane, and is hihgly conserved in all known oxygenic photosynthetic organisms. Alternative names of PsbO1 include 33 kDa subunit of oxygen evolving system of photosystem II, OEC 33 kDa subunit, 33 kDa thylakoid membrane protein, manganese-stabilizing protein 1 and for PsbO2 33 kDa subunit of oxygen evolving system of photosystem II, OEC 33 kDa subunit, 33 kDa thylakoid membrane protein, manganese-stabilizing protein 2.
- Product Citations
-
Selected references: Pipitone et al. (2021). A multifaceted analysis reveals two distinct phases of chloroplast biogenesis during de-etiolation in Arabidopsis. Elife. 2021 Feb 25;10:e62709. doi: 10.7554/eLife.62709. PMID: 33629953; PMCID: PMC7906606.
Pralon et al. (2019). Plastoquinone homoeostasis by Arabidopsis proton gradient regulation 6 is essential for photosynthetic efficiency. Commun Biol. 2019 Jun 20;2:220. doi: 10.1038/s42003-019-0477-4.
- Protocols
-
Agrisera Western Blot protocol and video tutorials
Protocols to work with plant and algal protein extracts
Agrisera Educational Posters Collection - Reviews:
-
This product doesn't have any reviews.