2
Anti-RbcS | Rubisco small subunit (SSU)
AS07 259 | Clonality: Polyclonal | Host: Rabbit | Reactivity: A. thaliana, C. pepo, C. reinhardtii, C. sativus, H. vulgare, M. domestica, N. tabacum, S. lycopersicum
- Product Info
-
Immunogen: KLH-conjugated synthetic peptide derived from all known sequences of RbcS from monocots and dicots including RuBisCO small subunit 1A UniProt: P10795,TAIR: AT1G67090, and 1B of Arabidopsis thaliana UniProt: P10796
At5g38430Host: Rabbit Clonality: Polyclonal Purity: Serum Format: Lyophilized Quantity: 50 µl Reconstitution: For reconstitution add 50 µl of sterile water Storage: Store lyophilized/reconstituted at -20°C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Western blot (WB) Recommended dilution: 1 : 5000 (WB) Expected | apparent MW: 20 | 15 kDa
- Reactivity
-
Confirmed reactivity: Arabidopsis thaliana, Brassica napus, Chlamydomonas reinhardtii, Cucumis sativus, Cucurbita pepo, Cyanthobasis fruticulosa, Hordeum vulgare, Malus domestica, Nicotiana tabacum, Petrosimonia nigdeensis, Salsola grandis, Salsola tragus, Solanum lycopersicum
Predicted reactivity: Algae, Camellia oleifera, Erythranthe guttata, Flaveria bidentis, Flaveria sonorensis, Glycine max, L, Marchantia paleacea, Musa acuminata, Nicotiana benthamiana, Nicotiana tabacum, Oryza sativa, Petunia hybrida, Polianthes tuberosa, Populus deltoides, Triticum aestivum, Solanum melongena, Solanum tuberosum, Zea mays
Species of your interest not listed? Contact usNot reactive in: Cyanobacteria - Application Examples
-
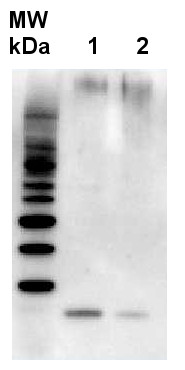
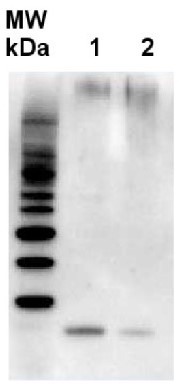
2 µg of total protein from Arabidopsis thaliana (1), Hordeum vulgare (2), extracted with Agrisera PEB extraction buffer (AS08 300) Samples were diluted with 1X sample buffer (NuPAGE LDS sample buffer (Invitrogen) supplemented with 50 mM DTT and heat at 70°C for 5 min and keept on ice before loading. Protein samples were separated on 4-12% Bolt Plus gels, LDS-PAGE and blotted for 70 minutes to PVDF using tank transfer. Blots were blocked immediately following transfer in 2% blocking reagent or 5% non-fat milk dissolved in 20 mM Tris, 137 mM sodium chloride pH 7.6 with 0.1% (v/v) Tween-20 (TBS-T) for 1h at room temperature with agitation. Blots were incubated in the primary antibody at a dilution of 1: 10 000 (in blocking reagent) for 1h at room temperature with agitation. The antibody solution was decanted and the blot was rinsed briefly twice, and then washed 1x15 min and 3x5 min with TBS-T at room temperature with agitation. Blots were incubated in secondary antibody (goat anti-rabbit IgG horse radish peroxidase conjugated, recommended secondary antibody AS09 602, Agrisera) diluted to 1:25 000 in blocking reagent for 1h at room temperature with agitation. The blots were washed as above. The blot was developed for 5 min with chemiluminescent detection reagent according the manufacturers instructions. Images of the blots were obtained using a CCD imager (VersaDoc MP 4000) and Quantity One software (Bio-Rad). Exposure time was 25 seconds.
Courtesy of Mayura Manerkar, Mount Alison University, Canada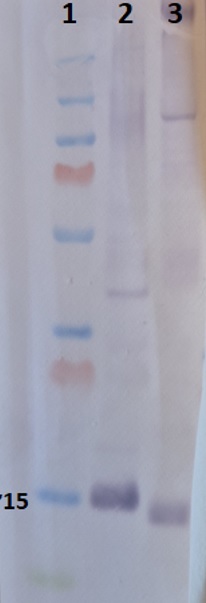
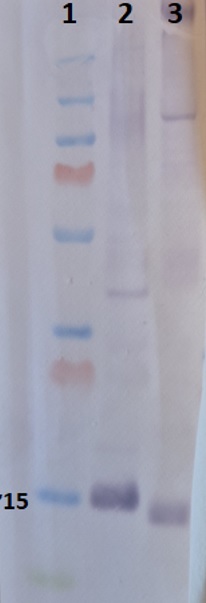
5 µg/well of total protein extracted freshly from Arabidopsis thaliana (2) and Zea Mays (3) denatured with 4XLDS at 70°C for 5 min. Protein were separated on NuPAGE Bis-Tris SDS gel and blotted 1h to Invitrogen PVDF (pore size of 0.2 um), using wet transfer. Blot was blocked with 5% milk 4°C/ON with agitation. Blot was incubated in the primary antibody at a dilution of 1: 1 000 for 1h/RT with agitation in TBS-T Blocking. The antibody solution was decanted, and the blot was rinsed briefly, then washed 3 times for 5 min in TBS-T at RT with agitation. Blot was incubated in Agrisera matching secondary antibody (AS09 607) diluted to 1:2500 in TBS-T Blocking for 30min/RT with agitation. The blot was washed as above and developed for 2 min with Agrisera BCIP/NBT plus. - Additional Information
-
Additional information (application): This product can be sold containing ProClin if requested - Background
-
Background: Rubisco catalyzes the rate-limiting step of CO2 fixation in photosynthesis. This enzyme contains two subunits, each present in eight copies. In plants and green algae, 55-kD large subunit is coded by the chloroplast rbcL gene, and the 15-kD small subunit is coded by a family of nuclear RbcS genes.
- Product Citations
-
Selected references: Kocacinar et aL. (2025). Biochemical and structural differences between C3 cotyledons and C4 leaves in species of Salsoloideae (Chenopodiaceae). Sci Rep. 2025 Oct 17;15(1):36383.doi: 10.1038/s41598-025-20388-w.
Lim et al (2022). Arabidopsis guard cell chloroplasts import cytosolic ATP for starch turnover and stomatal opening. Nat Commun. 2022 Feb 3;13(1):652. doi: 10.1038/s41467-022-28263-2. PMID: 35115512; PMCID: PMC8814037.
Mazur et al. (2021) The SnRK2.10 kinase mitigates the adverse effects of salinity by protecting photosynthetic machinery. Plant Physiol. 2021 Dec 4;187(4):2785-2802. doi: 10.1093/plphys/kiab438. PMID: 34632500; PMCID: PMC8644180.
Bernau et al. (2021) Precision analysis for the determination of steric mass action parameters using eight tobacco host cell proteins,Journal of Chromatography A,Volume 1652,2021,462379,ISSN 0021-9673,https://doi.org/10.1016/j.chroma.2021.462379. (https://www.sciencedirect.com/science/article/pii/S0021967321005033)
Ma et al. (2020). An ortholog of the Vasa intronic gene is required for small RNA-mediated translation repression in Chlamydomonas reinhardtii. Proc Natl Acad Sci U S A. 2020 Jan 7;117(1):761-770. doi: 10.1073/pnas.1908356117.
Akmouche et al. (2019). Do nitrogen- and sulphur-remobilization-related parameters measured at the onset of the reproductive stage provide early indicators to adjust N and S fertilization in oilseed rape (Brassica napus L.) grown under N- and/or S-limiting supplies? Planta. 2019 Dec;250(6):2047-2062. doi: 10.1007/s00425-019-03284-2.
Karimzadegan et al. (2018). The Effect of Methyl Jasmonate and Temperature on the Transient Expression of Recombinant Proteins in Cucurbita pepo L. Mol Biotechnol. 2018 Nov 27. doi: 10.1007/s12033-018-0138-8.
Luan et al. (2018). Elucidating the hypoxic stress response in barley (Hordeum vulgare L.) during waterlogging: A proteomics approach. Sci Rep. 2018 Jun 25;8(1):9655. doi: 10.1038/s41598-018-27726-1.
Shanmugabalaji et al. (2018). Chloroplast Biogenesis Controlled by DELLA-TOC159 Interaction in Early Plant Development. Curr Biol. 2018 Aug 20;28(16):2616-2623.e5. doi: 10.1016/j.cub.2018.06.006.
Ravi et al. (2018). Separation Options for Phosphorylated Osteopontin from Transgenic Microalgae Chlamydomonas reinhardtii. Int J Mol Sci. 2018 Feb 16;19(2). pii: E585. doi: 10.3390/ijms19020585.
Hartings et al. (2017). The DnaJ-Like Zinc-Finger Protein HCF222 Is Required for Thylakoid Membrane Biogenesis in Plants. Plant Physiol. 2017 Jul;174(3):1807-1824. doi: 10.1104/pp.17.00401.
Yin et al. (2016). Interplay between mitogen-activated protein kinase and nitric oxide in brassinosteroid-induced pesticide metabolism in Solanum lycopersicum. J Hazard Mater. 2016 Oct 5;316:221-31. doi: 10.1016/j.jhazmat.2016.04.070. Epub 2016 Apr 29.
Robert et al. (2015). Leaf proteome rebalancing in Nicotiana benthamiana for upstream enrichment of a transiently expressed recombinant protein. Plant Biotechnol J. 2015 Aug 19. doi: 10.1111/pbi.12452.
Dahal et al. (2015). Improved photosynthetic performance during severe drought in Nicotiana tabacum overexpressing a nonenergy conserving respiratory electron sink. New Phytol. 2015 May 29. doi: 10.1111/nph.13479.
Krasuska et al. (2015). Switch from heterotrophy to autotrophy of apple cotyledons depends on NO signal. Planta. 2015 Jul 18.
Huang et al. (2015). Rubisco accumulation is important for the greening of the fln2-4 mutant in Arabidopsis. Volume 236, July 2015, Pages 185–194.
Kim et al. (2015). Cytosolic targeting factor AKR2A captures chloroplast outer membrane-localized client proteins at the ribosome during translation. Nat Commun. 2015 Apr 16;6:6843. doi: 10.1038/ncomms7843.
Sun et al. (2014). The response of rbcL, rbcS and rca genes in cucumber, (Cucumis sativus L.) to growth and induction light intensity. Acta Physiol Plant, October 2014, Volume 36, Issue 10, pp 2779-2791 - Protocols
-
Agrisera Western Blot protocol and video tutorials
Protocols to work with plant and algal protein extracts
Oxygenic photosynthesis poster by prof. Govindjee and Dr. Shevela
Z-scheme of photosynthetic electron transport by prof. Govindjee and Dr. Björn and Dr. Shevela - Reviews:
-
Micha? Tracz | 2019-11-26Antibodies work well with the recommended dillution. My load per lane was an equivalent of total protein extracted from 1.5 mg leaf wet weight.Claus Schwechheimer | 2009-05-08Worked very nice on WB with 15µg of total proteins extracted from Arabidopsis thaliana.


