1
Anti-SMT1 | Sterol methyltransferase 1
AS07 266 | Clonality: Polyclonal | Host: Rabbit | Reactivity: Arabidopsis thaliana | Marker antibody of integral ER membrane
- Product Info
-
Immunogen: KLH-conjugated synthetic peptide derived from Arabidopsis thaliana SMT1 UniProt: Q9LM02, TAIR: At5g13710
Host: Rabbit Clonality: Polyclonal Purity: Immunogen affinity purified serum in PBS pH 7.4. Format: Lyophilized Quantity: 100 µg Reconstitution: For reconstitution add 100 µl of sterile water Storage: Store lyophilized/reconstituted at -20C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Immunolocalization (IL), Western blot (WB) Recommended dilution: 1 : 50-1 : 100 (IL), 1 : 500-1 : 1000 (WB) Expected | apparent MW: 38 kDa
- Reactivity
-
Confirmed reactivity: Arabidopsis thaliana Predicted reactivity: Amborella trichopoda, Brassica napus, Brassica rapa, Capsella rubella, Citrus clementina, Eutrema salsugineum, Glycine max, Glycine soja, Gossypium mexicanum, Medicago truncatula, Populus trichopocarpa, Prunus persica, Ricinus communis, Theobroma cacao, Vitis vinifera
Species of your interest not listed? Contact usNot reactive in: Hordeum vulgare, Triticum aestivum, Solanum lycopersicum, Withania somnifera
- Application Examples
- Application example
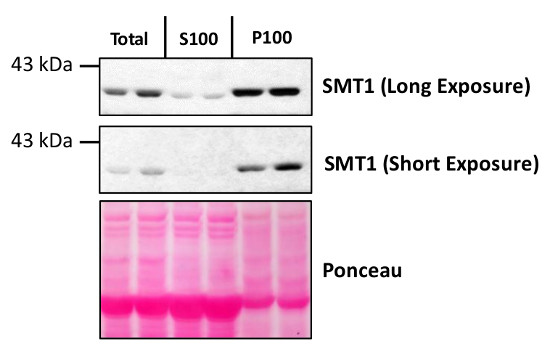
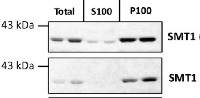
Total protein from Col-0 (wild-type) Arabidopsis thaliana were extracted with 50mM HEPES-KOH buffer containing 250 mM sucrose, 5% glycerol, 50 mM NaPP, 1 mM NaMo, 25 mM NaF, 10mM EDTA, 0.5% PVP, 3mM DTT, 1mM PMSF, 10uM Leupeptin & 10nM Calyculin, and then fractionated by ultracentrifugation at 100,000 x gravity for 30 min at 4°C into soluble (S100) and microsomal (P100) proteins as described in LaMontagne et al. (2016). Isolation of Microsomal Membrane Proteins from Arabidopsis thaliana. Current Protocols in Plant Biology 1:1-18. doi: 10.1002/cppb.20020. 30 µg proteins of total, S100 and P100 fractions were denatured at 37°C for 5 min, separated on a 7.5 % SDS-PAGE and blotted 1h to nitrocellulose using tank transfer. Blots were blocked with 1x PBS (from Fisher Scientific BP665-1) + 0.1 %Tween 20 (PBS-T) + 5% milk for 1h at room temperature (RT) with agitation. Blot was incubated in the primary antibody at a dilution of 1: 1000 overnight at 4°C with agitation in 1x PBS-T + 5% milk. The antibody solution was decanted and the blot was rinsed briefly once, then washed four times for 7 min in 1x PBS-T at RT with agitation. Blot was incubated in secondary antibody (anti-rabbit IgG horse radish peroxidase conjugated) diluted to 1:10 000 in 1x PBS-T + 5% milk for 2 hrs at RT with agitation. The blot was washed as above and developed for 4 min with chemiluminescent detection reagent. Exposure time was 30 seconds and 2 min.Courtesy of Erica LaMontagne & Dr. Antje Heese (Division of Biochemistry, Interdisciplinary Plant Group (IPG) - University of Missouri; Columbia, MO, USA)Application examples: 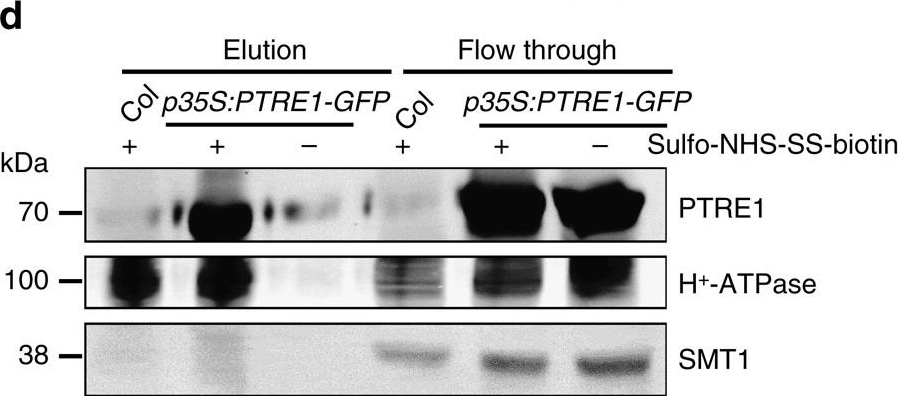
Reactant: Arabidopsis thaliana (Thale cress)
Application: Western Blotting
Pudmed ID: 27109828
Journal: Nat Commun
Figure Number: 1D
Published Date: 2016-04-25
First Author: Yang, B. J., Han, X. X., et al.
Impact Factor: 13.783
Open PublicationProtein structure and subcellular localization of PTRE1.(a) PTRE1 has conserved FP and proline-rich domains similar to human PI31, plant-specific domains and a N-terminal transmembrane region (upper panel). An amino acid alignment of the C-terminal proline-rich domain, which is conserved in the predicted proteasome inhibitor family of eukaryotic species (hPI31, Homo sapiens; LOC682071, Rattus norvegicus; and Os07g054890, Oryza sativa), is shown (bottom panel). The R(Ar)DP motif is indicated by the purple rectangle and the conserved amino acid residues of these four proteins are highlighted with different colours. (b) PTRE1 localizes at nucleus (upper; scale bar, 20??m), plasma membrane (middle; bar, 20??m) and ER (bottom; bar, 20??m). Pavement cells of Arabidopsis seedlings expressing PTRE1-GFP were stained with DAPI (2??g?ml?1) to show the nuclear localization of PTRE1 (upper, DAPI stains nucleus and plant cell walls). Hypocotyl cells of 7-day-old Arabidopsis seedlings expressing PTRE1-GFP and plasma membrane aquaporin PIP2-RFP was observed after plasmolysis (middle). Plasmolysis was performed by adding 0.6?M mannitol for 30?min. PTRE1-GFP was transiently expressed in N. benthamiana leaves with ER-mCherry (bottom) and observed. (c) Enlarged ER localization of PTRE1 by transient expression of PTRE1-GFP or ER-mCherry in N. benthamiana leaves. Scale bar, 20??m. (d) Parts of PTRE1 protein were localized and surface-exposed at the plasma membrane. Plant materials were treated with the membrane-impermeable sulpho-NHS-SS-biotin reagent (+). Surface-exposed protein eluted after purification using biotin beads. Plasma membrane protein H+-ATPase and ER membrane protein SMT1 were used as control. Equal amounts of samples were subjected to SDS–PAGE and immunoblot analysed using anti-PTRE1 (rabbit), anti-H+-ATPase (rabbit) or anti-SMT1 (rabbit) antibodies.
- Additional Information
-
Additional information: SMT1 is an integral membrane protein of ER (Boutte et al., 2009) while BiP is a membrane associated protein.
Endogenous SMT1 is expressed at very low levels in root epidermis and root cap as compare to cortex and endodermis and therefore this can contribute to SMT1 detection problems.Additional information (application): SMT1 antibody characterization in Western Blot and immunofluorescence labeling: Boutté Y et al. (2010). Endocytosis restricts Arabidopsis KNOLLE syntaxin to the cell division plane during late cytokinesis. EMBO J. 29, 5465-5458. - Background
-
Background: SMT1 (EC=2.1.1.41) is an enzyme involved in steroid biosynthesis localized specific to the Endoplasmic Reticulum (ER), Boutte et al., 2009 . It is catalyzing the methyl transfer from S-adenosyl-methionine to the C-24 of cycloartenol to form 24-methylene cycloartenol. The protein is highly expressed in vascular tissue, mature leaves and in regions undergoing cellular expansion. Alternative names: Cycloartenol-C-24-methyltransferase, 24-sterol C-methyltransferase 1, Protein STEROL METHYLTRANSFERASE 1, Protein CEPHALOPOD
- Product Citations
-
Selected references: Ming-fang et al. (2021) Improved quantification of immune-gold labeling and its use to compare the distribution of cellular factors among sub-chloroplast compartments,Micron,2021,103060,ISSN 0968-4328,https://doi.org/10.1016/j.micron.2021.103060.
Cano-Ramirez et al. (2021) M. Plasma Membrane Fluidity: An Environment Thermal Detector in Plants. Cells. 2021 Oct 17;10(10):2778. doi: 10.3390/cells10102778. PMID: 34685758; PMCID: PMC8535034.
Collins et al. (2020). EPSIN1 Modulates the Plasma Membrane Abundance of FLAGELLIN SENSING2 for Effective Immune Responses . Plant Physiol. 2020 Feb 24. pii: pp.01172.2019. doi: 10.1104/pp.19.01172
Laohavisit (2020). Quinone perception in plants via leucine-rich-repeat receptor-like kinases. Nature. 2020 Nov;587(7832):92-97. doi: 10.1038/s41586-020-2655-4. Epub 2020 Sep 2. PMID: 32879491.
Yang et al. (2016). Arabidopsis PROTEASOME REGULATOR1 is required for auxin-mediated suppression of proteasome activity and regulates auxin signalling. Nat Commun. 2016 Apr 25;7:11388. doi: 10.1038/ncomms11388.
LaMontagne et al. (2016). Isolation of Microsomal Membrane Proteins from Arabidopsis thaliana. Curr. Protoc. Plant Biol. 1:217-234. doi: 10.1002/cppb.20020.
Yoshimoto et al. (2014). Quality control of plant peroxisomes in organ specific manner via autophagy. J cell Science, August 1, 2014, 127 (15). - Protocols
-
Agrisera Western Blot protocol and video tutorials
Protocols to work with plant and algal protein extracts - Reviews:
-
This product doesn't have any reviews.
Related products
AS09 481 | Clonality: Polyclonal | Host: Rabbit | Reactivity: A. thaliana, B. napus, Chara australis, C. reinhardtii, C. sativus, M. perniciosa, N. benthamiana, N. tabacum, R. sativa, L.Tokinashi-daikon, O. europaea, O. sativa, P. abies, P. patens, S. oleracea, S. lycopersicum, S. tuberosum, T. aestivum, Z. mays


