1
Anti-Cyt c | Cytochrome c
AS08 343A | Clonality: Polyclonal | Host: Rabbit | Reactivity: A. thaliana, B. oleracea, G. max, L. luteus, P. satiovum, Z. mays | Marker of PCD
- Product Info
-
Immunogen: KLH-conjugated synthetic peptide derived from Arabidopsis thaliana cytochrome c protein sequence, UniProt:D7KMK0 (C-1) D7LY03 (C-2), TAIR: At1g22840 (Cytc1) and At4g10040 (Cytc2)
Host: Rabbit Clonality: Polyclonal Purity: Immunogen affinity purified serum in PBS pH 7.4. Format: Lyophilized Quantity: 50 µg Reconstitution: For reconstitution add 50 µl of sterile water Storage: Store lyophilized at -20°C; once reconstituted make aliquots to avoid repeated freeze-thaw cycles and Store at -80°C. Please remember to spin the tubes briefly prior to opening them to avoid any losses that might occur from material adhering to the cap or sides of the tube. Tested applications: Immunolocalization (IL), Western blot (WB) Recommended dilution: 1: 100 (IL), 1 : 5000 (WB) Expected | apparent MW: 12.5 | 14 kDa (for Arabidopsis thaliana)
- Reactivity
-
Confirmed reactivity: Arabidopsis thaliana, Brassica oleracea, Glycine max, Pisum sativum, Zea mays
Predicted reactivity: cytc1 and cytc2 from following species: A. theoprasi, Brassica napus, Brassica oleracea, Cannabis sativa, C. maxima, Chlamydomonas reinhardtii (peptide target partially conserved), Lupinus luteus, Medicago truncatula, Nicotiana tabacum, Oryza sativa, Ostreococcus (peptide target partially conserved), P. aurea, Physcomitrium patens, Ricinus communis, S. nigra, Solanum lycopersivum, Vigna radiata, Vitis vinifera.
Species of your interest not listed? Contact usNot reactive in: Arabidopsis thaliana CytC6, Chinesecuscuta sp. - Application Examples
-
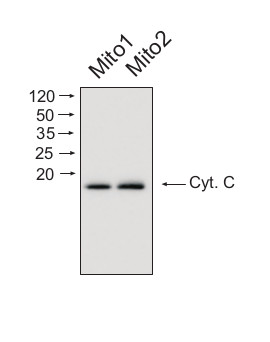
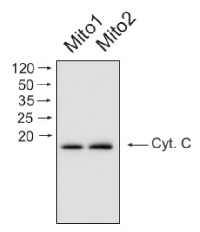
Mitochondrial proteins (15 ug) from Arabidopsis thaliana mitochondria was separated on 16% acrilamide gel and electrophoresis prepared according to Schägger and von Jagov (Anl. Biochem., 1987, 166:368-379). After running the gel, proteins were transferred to PVDF membrane using wet transfer (Roti®-Blot 2, Roth). Transfer was checked by Ponceau S staining. Blot was destained by several quick washings in distilled water and 1 washing in 1X TBS (10 mM T pH 7.5, 150 mM NaCl) (10-15 min.).Blot was blocked by 1.5 hour in 5% milk in TBST (1X TBS, 0,1% Tween 20) After blocking blot was washed quickly twice in TBST and incubated 2 hours with primary antibody (dilution 1: 1000) in TBST. Washing: two quick washings in TBST and 3 x 10 min. washings in TBST. Then blot was incubated 45-60 min. with a secondary anti-rabbit antibodies conjugated to peroxidase (Agrisera AB, dilution 1:10 000, AS09 602) in TBST. Washing: as above. After washing blot was incubated 1-2 min. in chemiluminescent detection reagent. Chemiluminescence was detected by BioSpectrum® Imaging System (UVP). Exposure time was 5 seconds.
Courtesy Dr. Janusz Piechota, Wrocław University, Poland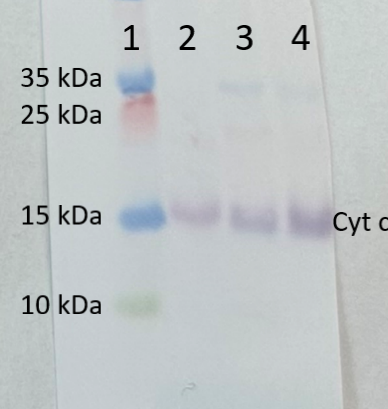
Lane:
1 - MW marker
2 - 10 ug of Arabidopsis thaliana whole leaf extract
3 - 10 ug of Zea mays whole leaf extract
4- 10 ug of Solanum lycopersicum whole leaf extract
10 µg/well of total protein extracted from Arabidopsis thaliana, Zea mays and Solanum lycopersicum, total cell extract, stored at -80°C. Exact buffer components were 1xPEB (AS08 300) with protease inhibitor (0.1 mg/ml for Pefabloc SC, Roche) and denatured with Invitrogen LDS sample buffer (4X) at 70°C/5 min. Samples were separated on Invitrogen NuPage Bis-Tris 4-12% SDS-PAGE and blotted for 1 h to Invitrogen PVDF (pore size of 0.2 um), using wet transfer. Blot was blocked with 5% milk in TBS-T for 1h/RT with agitation. Blot was incubated in the primary antibody at a dilution of 1: 500 for 1h/RT with TBS-T Blocking. The antibody solution was decanted, and the blot was rinsed briefly twice, then washed once for 15 min and 3 times for 5 min in TBS-T at RT with agitation. Blot was incubated in matching secondary antibody (anti-rabbit IgG ALP conjugated, AS09 607 lot 2302 ) diluted to 1: 1 000 in TBS-T Blocking for 0,5h/RT with agitation. The blot was washed as above and developed with AS19 BCIP-NBT-PLUS lot 09269221 for 3min. As soon as the desired band was detectable, the membrane was washed in generous amounts of deionized water before placing the membrane on Whatman paper to dry. Image was captured after 1 h.
Courtesy Agrisera - Additional Information
-
Additional information (application): https://www.agrisera.com/en/artiklar/goat-anti-rabbit-igg-hl-ap-conjugated-.htmlThe presence of cytochrome c in the cysotol is a marker of PCD (programmed cell death) - Background
-
Background: Cytochrome c is located in inner mitochondrial membrane. It is a small heme protein which, unlike other cytochromes, is highly soluble. This protein is an essential component of the electron transport chain, where it undergoes oxidation and reduction without binding oxygen.
- Product Citations
-
Selected references: Law et al. (2025). Mitochondria-Specific Protein Delivery by Protease-Triggered Release in Plants with Single-Walled Carbon Nanotubes. Advanced Materials Technologies, doi.org/10.1002/admt.202501468.
Soria et al. (2024).Functional resilience: An active oxidative phosphorylation system prevails amid foreign proteins in holoparasitic plants. Current Plant Biology Volume 37, March 2024, 100322.
Canal et al. (2024).Cytochrome c levels affect the TOR pathway to regulate growth and metabolism under energy-deficient conditions. New Phytol. 2024 Mar;241(5):2039-2058. Guo et al. (2021) The pentatricopeptide repeat protein GEND1 is required for root development and high temperature tolerance in Arabidopsis thaliana,Biochemical and Biophysical Research Communications,Volume 578,2021,Pages 63-69,ISSN 0006-291X,https://doi.org/10.1016/j.bbrc.2021.09.022.(https://www.sciencedirect.com/science/article/pii/S0006291X21013164)
Wang et al. (2020) Rerouting of ribosomal proteins into splicing in plant organelles. BioRxiv, DOI: 10.1101/2020.03.03.974766 .
Dai et al. (2020). Pentatricopeptide repeat protein DEK46 is required for multi-sites mitochondrial RNA editing and maize seed development. J Exp Bot. 2020 Jul 25;eraa348.doi: 10.1093/jxb/eraa348.
Waltz et al. (2019). Small is big in Arabidopsis mitochondrial ribosome. Nat Plants. 2019 Jan;5(1):106-117. doi: 10.1038/s41477-018-0339-y.
Doronina et al. (2019). Structural and Functional Features of the Wheat Embryo Sac’s Antipodal Cells during Differentiation. Russ J Dev Biol 50, 194–208. (immunolocalization)
Waltz et al. (2019). Small is big in Arabidopsis mitochondrial ribosome. Nat Plants. 2019 Jan;5(1):106-117. doi: 10.1038/s41477-018-0339-y.
Doronina et al. (2019). Structural and Functional Features of the Wheat Embryo Sac’s Antipodal Cells during Differentiation. Russ J Dev Biol 50, 194–208. (immunolocalization)
Waltz et al. (2019). Small is big in Arabidopsis mitochondrial ribosome. Nat Plants. 2019 Jan;5(1):106-117. doi: 10.1038/s41477-018-0339-y.
Doronina et al. (2019). Structural and Functional Features of the Wheat Embryo Sac’s Antipodal Cells during Differentiation. Russ J Dev Biol 50, 194–208. (immunolocalization).
Rurek et al. (2018). Mitochondrial Biogenesis in Diverse Cauliflower Cultivars under Mild and Severe Drought Involves Impaired Coordination of Transcriptomic and Proteomic Response and Regulation of Various Multifunctional Proteins. Preprints 2018, 2018010276 (doi: 10.20944/preprints201801.0276.v1).
Dai et al. (2018). Maize Dek37 Encodes a P-Type PPR Protein That Affects Cis-splicing of Mitochondrial nad2 Intron 1 and Seed Development. Genetics. 2018 Jan 4. pii: genetics.300602.2017. doi: 10.1534/genetics.117.300602.
Opalinska et al. (2017). Identification of Physiological Substrates and Binding Partners of the Plant Mitochondrial Protease FTSH4 by the Trapping Approach. Int J Mol Sci. 2017 Nov 18;18(11). pii: E2455. doi: 10.3390/ijms18112455.
Schimmeyer et al. (2016). L-Galactono-1,4-lactone dehydrogenase is an assembly factor of the membrane arm of mitochondrial complex I in Arabidopsis. Plant Mol Biol. 2016 Jan;90(1-2):117-26. doi: 10.1007/s11103-015-0400-4. Epub 2015 Oct 31.
Li et al. (2016). Characterization of a novel Beta-barrel protein (AtOM47) from the mitochondrial outer membrane of Arabidopsis thaliana. J Exp Bot. 2016 Nov;67(21):6061-6075. Epub 2016 Oct 6. - Protocols
-
Agrisera Western Blot protocol and video tutorials
Protocols to work with plant and algal protein extracts
Oxygenic photosynthesis poster by prof. Govindjee and Dr. Shevela
Z-scheme of photosynthetic electron transport by prof. Govindjee and Dr. Björn and Dr. Shevela - Reviews:
-
Elisa | 2018-08-23Very good detection with colorimetric reaction on Western blotting with mitochondrial proteins from soybean and pea plant tissuesAdrian | 2017-04-27Worked very well for western-blotting in the lace plant (Aponogeton madagascariensis)
Related products
AS04 053A | Clonality: Polyclonal | Host: Rabbit | Reactivity: Eudicots, monocots, Physcomitrella patens | Cellular [compartment marker] of mitochondrial inner membrane


