1

Anti-PsbA | D1 protein of PSII, C-terminal, HRP-conjugated (40 µg)
AS05 084A-HRP | Clonality: Polyclonal | Host: Rabbit | Reactivity: [global antibody] for higher plants, algae, liverwort, cyanobacteria, diatoms compartment marker of thylakoid membrane
Benefits of using this antibody
- Product Info
-
Immunogen: KLH-conjugated synthetic peptide derived from available plant, algal and cyanobacterial PsbA sequences, including Arabidopsis thaliana UniProt: A4QJR4, TAIR: AtCg00020 , Oryza sativa P0C434, Populus alba Q14FH6, Physcomitrella patens Q6YXN7, Chlamydomonas reinhardtii P07753, Synechocystis sp. P14660 and many others
Host: Rabbit Clonality: Polyclonal Purity: Immunogen affinity purified serum in PBS pH 7.4, conjugated to HRP. Format: Liquid Quantity: 40 µg Storage: Store at 4°C for 12-18 months. A preservative may be added for long time storage up to 2 years. Tested applications: Western blot (WB) Recommended dilution: 1 : 15 000 (WB) Expected | apparent MW: 38 | 28-30 kDa
- Reactivity
-
Confirmed reactivity: Anabaena 7120, Arabidopsis thaliana, Artemisia annua, Arundo sp., Chlamydomonas reinhardtii, Colobanthus quitensis Kunt Bartl, Coscinodiscus wailesii, Craterostigma sp., Ditylum brightwellii, Glycine max, Hordeum vulgare, Lindernia sp., Miscanthus x giganteus, Marchantia polymorpha (liverwort), Nicotiana benthamiana, Panicum miliaceum, Panax ginseng, Panicum maximum, Pinus strobus, Physcomitrium patens, Synechococcus sp. PCC 7942, Paulinella chromatophora (amoeba), Prochlorococcus sp. (surface and deep water ecotype), Spirodela polyrhiza, Symbiodinium sp., Zea mays Predicted reactivity: Algae (brown and red), Brassica napus, Conifers, Cyanobacteria, Dictos, Cannabis sativa, Galdieria sulphuraria, Lactuca sativa, Lycopersicum esculentum, Medicago sativa, Nannochloropsis sp., Oryza sativa, Ostreococcus sp. Pisum sativum, Porphyridium purpureum, Sesamum indicum, Thalassiosira pseudonana, Zosteria marina, Vitis vinifera cellular [compartment marker] of thylakoid membrane
Species of your interest not listed? Contact usNot reactive in: No confirmed exceptions from predicted reactivity are currently known - Additional Information
-
Additional information: Due to biology of PsbA (D1) protein a number of degradation products can apprear in a sample and may be observed when using anti-PsbA antibodies, including products having apparent molecular weights of 24kDa and 16kDa. D1 degradation is a complex set of events and the products observed can be influenced by both the extraction procedure and the physiology of the cells prior to harvest. Third, cross-linking may occur between D1 and cytochrome b559, shifting the protein higher in the gel. In cyanobacteria (PCC7942), three different bands were competed out by preincubating the antibody with the PsbA free peptide, indicating that all bands are indeed PsbA and its precursors or breakdown products. Competition assays were also performed with spinach and Chlamydomonas, confirming the identity of PsbA bands.
Anti-PsbA antibodies will not detect D2 protein, as the peptide used to generate PsbA antibodies has no homology to the D2 sequence.
Additional information (application): The antibody is appropriate for detecting both, 24 kDa or the 10 kDa C-terminal fragments, whichever is generated under given treatment conditions. In our analysis we have seen both, ca. 24 kDa and ca. 10 kDa fragments from different samples, depending on treatments and isolation procedures.
Rabbit anti-PsbA antibody can detect more than one band of PsbA protein, e.g. precursor and mature protein as compare to the hen anti-PsbA antibodies AS01 016.
This antibody will detect the phosphorylated form of D1 as an alternate band to the main band on a high resolution gel.
The antibody will bind to cross-linked proteins: D1/D2, D1/cyt b559, D1/CP43. - Background
-
Background: The psbA gene has been cloned from many species of plants, green algae, and cyanobacteria. The psbA gene is located in the chloroplast genome and encodes for the D1 protein, a core component of Photosystem II. PsbA/D1 is rapidly cycled under illumination in all oxygenic photobionts. Tracking PsbA pools using the Global PsbA antibody can show the functional content of Photosystem II in a wide range of samples. Alternative names: 32 kDa thylakoid membrane protein, photosystem II protein D1. - Product Citations
-
Selected references: Thurotte et al. (2020). DnaK3 Is Involved in Biogenesis and/or Maintenance of Thylakoid Membrane Protein Complexes in the Cyanobacterium Synechocystis Sp. PCC 6803. Life (Basel). 2020 Apr 30;10(5):E55. doi: 10.3390/life10050055. - Protocols
-
Agrisera Western Blot protocol and video tutorials
Protocols to work with plant and algal protein extracts
PsbA quantitation in plant and algal samples using Agrisera anti-PsbA antibody and PsbA protein standard
Methodology: Plant samples are generally ground with liquid nitrogen in a mortar and pestle. The resulting powder is transferred to a plastic tube. Algal samples can be either concentrated by centrifugation or, preferably, by filtration onto glass fiber filters. Solubilization is performed in Agrisera protein extraction buffer (PEB, AS08 300) containing 0.1mg/mL PefaBloc SC (AEBSF) protease inhibitor (Roche). Disruption is most optimally obtained through flash freezing of the sample in liquid nitrogen alternated with thawing by sonication with a microtip. This process can be repeated depending on the toughness of the sample. The sample is adjusted to 50 mM dithiothreitol and heated to 70°C for 5 minutes. Samples are cooled and centrifuged briefly prior to electrophoresis.
Optimal quantitation is achieved using moderate sample loads per gel lane, generally 0.5 to 2.5 ug total protein, depending on the abundance of the target protein.
Electrophoresis and Immunoblotting: Once solubilized, the proteins can be separated electrophoretically in a number of systems. We obtain optimal results with the Invitrogen NuPAGE gel system using Bis-Tris 4-12% gradient gels. Proteins are separated in MES SDS running buffer according to the manufacturer’s recommendations at 200 V for 35 minutes. The gels are transferred to PVDF in the same apparatus, the SureLock XCell blot module, for 60 minutes at 30 V for a single gel or 80 minutes for a pair. Following transfer the blots are blocked with non-fat dry milk up to 10 % in TBS-T, for 1 h/RT with gentle agitation. The blot is incubated with primary antibody, usually at 1:25 000 to 1:50 000, for 1 h/RT (if extreme femtogram detection reagents are used) or in lower primary antibody dilution for less sensitivie reagents (mid picogram and lower).
For quantitation a relatively high primary antibody: target protein ratio gives more reliable results than immunoblots at low ratios of primary antibody:target protein.
The blot is washed extensively in TBS-T (twice briefly, once for 15 minutes and three times for five minutes). The blot is incubated with secondary antibody, for example goat anti-rabbit IgG horse radish peroxidase conjugated, AS09 602 (Agrisera), for 1h/RT. The blot is washed as above and developed with ECL detection reagents.Quantitation: When quantitated standards are included on the blot, the samples can be quantitated using the available software. Excellent quantitation can be obtained with images captured on the Bio-Rad Fluor-S-Max or equivalent instrument using Bio-Rad QuantityOne software. The contour tool is used to select the area for quantitation and the values are background subtracted to give an adjusted volume in counts for each standard and sample. Using above protocol linear standard curves are generated over 1-1.5 orders of magnitude range in target load. It is important to note that immunodetections usually show a strongly sigmoidal signal to load response curve, with a region of trace detection of low loads, a pseudolinear range and a region of saturated response with high loads. For immunoquantitation it is critical that the target proteins in the samples and the standard curve fall within the pseudolinear range. Our total detection range using this protocol spans over 2 orders of magnitude, but the quantifiable range is narrower.
Application example
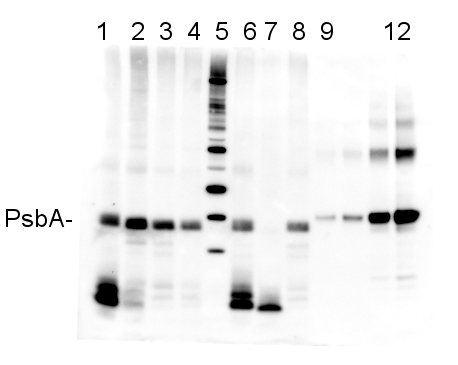
2 µg of total protein from (1-4 and 6-8) Prochlorococcus sp. extracted with Protein Extration Buffer, PEB (AS08 300), (9-12) PsbA protein standard (AS01 016S), (5) molecular weight markers (Magic Marks, Invitrogen) were separated on 4-12% NuPage (Invitrogen) LDS-PAGE and blotted 1h to PVDF. Blots were blocked immediately following transfer in 2% blocking reagent in Tween-20 (TBS-T) for 1h/RT with agitation. Blots were incubated in the primary antibody at a dilution of 1: 50 000 for 1h/RT with agitation. The antibody solution was decanted and the blot was rinsed briefly twice, then washed once for 15 min and 3 times for 5 min in TBS-T at room temperature with agitation. Blots were incubated in secondary antibody (anti-rabbit IgG horse radish peroxidase conjugated) diluted to 1:10 000 in for 1h/RT with agitation. The blots were washed as above and developed for 5 min with ECL detection reagent according the manufacturers instructions. Images of the blots were obtained using a CCD imager (FluorSMax, Bio-Rad) and Quantity One software (Bio-Rad). The nati Various smaller C-terminal cleavage products from PsbA are evident in the Prochlorococcus samples; these are real, and vary in reprocible ways depending on treatment and growth conditions.
Recommended secondary antibodies: goat anti-rabbit HRP conjugated, goat anti-rabbit ALP conjugated
Recommended chemiluminescent detection reagent: AgriseraECLBright
References:MacKenzie et al (2005). Large reallocations of carbon, nitrogen and photosynthetic reductant among phycobilisomes, photosystems and Rubisco during light acclimation in Synechococcus elongatus are constrained in cells under low environmental inorganic carbon. Arch of Microbiol. 183: 190 - 202.
Bouchard et al. (2006) UVB effects on the photosystem II-D1 protein of phytoplankton and natural phytoplankton communities. Photochem and Photobiol 82: 936-951. - Reviews:
-
This product doesn't have any reviews.
Related products
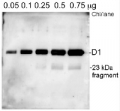
AS10 704 | Clonality: Polyclonal | Host: Rabbit | Reactivity: A.thaliana, C.reinhardii, H.vulgare, M.truncatula, N. oceanica, N. oleoabundans UTEX 1185 (chlorophyta), P.patens, P.sativum, S.vulgaris, S.alba, S.oleracea, Synechococcus sp. PCC 7942, Synechocystis sp. PCC6803, Triticum sp.
AS13 2669 | Clonality: Polyclonal | Host: Rabbit | Reactivity: [global antibody] for higher plants
Benefits of using this antibody